The Crystal Whisperer
Sarah Bowman
Hauptman-Woodward Medical Research Institute
Published November 29, 2023
In structural biology. there are few more onerous tasks than coaxing reluctant proteins into crystals. “Unfortunately, that is really a huge bottleneck,” says Sarah E.J. Bowman, who heads the National High-Throughput Crystallization (HTX) Center in upstate New York. The center aims to increase the odds of success for this essential step in much of structural biology.
Sooner or later, by old-fashioned trial and error, many researchers find the magic recipe to crystallize their protein at their own lab benches. After all, nearly 90% of structures deposited in the Protein Data Bank (PDB) started as crystals of sufficient quality to collect X-ray diffraction data. The resulting three-dimensional molecular models are essential tools to understand how proteins work and what goes wrong in diseases and to develop new vaccines and therapeutics.
But in the last two decades, some 2000 laboratories in academia, industry and elsewhere have opted in for the center’s screening service, sending more than 18,000 samples through (Lynch et.al, Acta Crystallographica Section D, 2023).
“We have a high-throughput pipeline—we're the only facility in the world that actually has this ability to do 1,536 different conditions in one experimental assay,” Bowman told an online audience. That adds up to crystallization screening data on more than 27 million screening experiments.
For the initial conditions screen, the center has mined its own records as well as PDB data to come up with chemical cocktails that have worked to crystallize proteins. The concoctions are a mix of in-house formulas, commercial solutions, and in-house adaptations. Crystal growth is tracked for 6 weeks.
The center’s robotic equipment deposits miniscule amounts of protein into 1,536 tiny oil-coated wells on a microarray plate designed either for soluble or membrane proteins. For scale, the diameter of a U.S penny would span 8 of the 1,536 tiny wells across in the microassay plate.
The frequent imaging (yielding about 14,000 images per sample) to track incremental progress of newborn crystals puts even the most doting new parents to shame. The center deploys several advanced imaging techniques to detect protein crystals and distinguish them from salt crystals or other substances. A post-baccalaureate student developed a graphical user interface, called MARCO Polo, to apply a machine learning algorithm to classify the crystals by (Holleman et.al, Journal of Applied Crystallography, 2021). Then the center can help optimize crystallization for reproducible results.
In early 2020, when the COVID-19 pandemic was declared, the center pivoted to focus exclusively on SARS-CoV-2 viral proteins, sometimes in complex with a potential drug. Over 2 years, thanks to dedicated funding from the National Science Foundation, the center’s users deposited more than 30 viral protein structures in the PDB, including the main protease in contact with inhibitors, and structures are still being deposited.
“Structural work has been absolutely critical for the development of both the vaccines and therapeutics or drug treatments against SARS-CoV-2,” Bowman notes. “It's been a really fascinating time to be a scientist, because so many different kinds of scientists came together and really worked together and communicated things very openly and quickly. And I think that ultimately has helped with generating treatments and vaccines.”
The center’s capabilities are evolving in concert with new experimental and measurement technologies. Even as cryo-electron microscopy is enabling structural solutions of ever-larger molecules, crystallography is getting smaller. Crystals previously considered too tiny to use may now provide valuable data, thanks to dedicated microfocus beamlines at synchrotron radiation sources and other new technologies and techniques.
“When I started, we were in the stage of ‘Make your crystals as big and chunky as you possibly can and collect all your data on one crystal.’ It’s still totally a great thing to do, but that has shifted in past 10 years,” says Bowman, who distinguishes macromolecular crystallography from micro techniques.
In one example of using very small crystals for structural work, in 2022, an international team of researchers reported the structure and mechanism of action of a newly identified antibiotic (Miller et.al, Nature Microbiology, 2022).
It was one of the first examples of the center’s three-way imaging to detect microcrystals and nanocrystals to use in microcrystal electron diffraction (microED), a technique that uses cryo-em instruments, Bowman says. For the study, Bowman helped collaborators at Northeastern University who had isolated the natural product from a computational genomic screen of bacteria that live in certain roundworms and help them parasitize insect larvae.
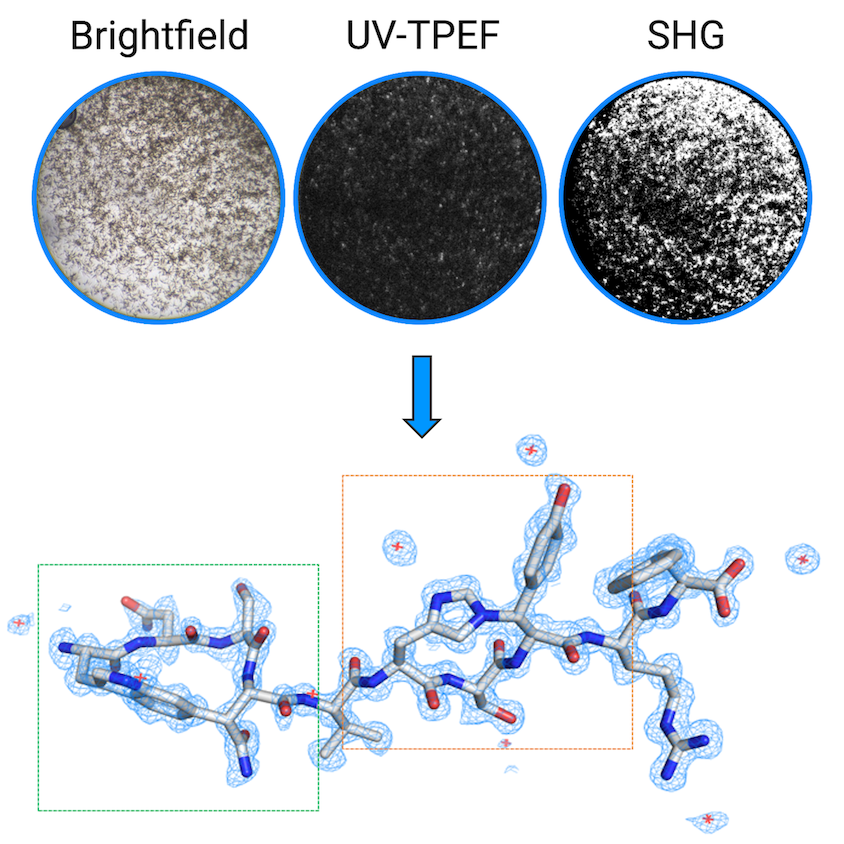

Bowman splits her time between running the center and her own lab. “One of our research projects is to develop better ways to handle these really small crystals, because it turns out that, just like in normal crystallography, this is actually one of the big bottlenecks.”
Questions include: How do you even know if you have a really small crystal? And: What is the best data-gathering technique based on the size of the crystal? “It turns out there is a huge window of opportunity for things [such as very small crystals] that were previously intractable,” Bowman says.
“I get very excited about the kind of cutting-edge things that are happening—new technology, detectors, the way we handle samples, even data processing—that are still in development and being pushed forward,” she says.

Bowman wasn’t always as excited about science. In fact, “I actually really disliked science,” she says. “In my first bachelor's degree—English lit and women’s studies [Cornell College, Iowa]—I don't think I took a single science class.” She considered going on to graduate school in English. She loved books, but she decided she just wanted to read them not dissect them.
Instead, she moved to Denver and joined the management team at the independent bookstore the Tattered Cover. In her human resources duties, she noticed many of the employees she was training were taking college classes in science they enjoyed.
So she sampled a few chemistry classes. Bowman had “an amazing professor who opened my eyes to science” and soon embarked on a second undergraduate degree at Metropolitan State College of Denver. She graduated with a BS in chemistry in 2005. One summer, she took a leave of absence from the bookstore to work fulltime in a laboratory. “It was wonderful,” she says. “I love being in the lab.”
She moved to upstate New York as a graduate student in chemistry at University of Rochester. In the lab of Kara Bren, Bowman applied nuclear magnetic resonance spectroscopy to metalloproteins to study their behavior. She visited a synchrotron for the first time while a graduate student, not for structural studies but to use a specialized spectroscopic technique called nuclear resonance vibrational spectroscopy. She became entranced with the experimental possibilities available at synchrotrons.
For her postdoctoral work, Bowman joined the labs of Catherine Drennan and Collin Stultz at Massachusetts Institute of Technology. It was at MIT that she pivoted toward structural biology, learning X-ray crystallography and computational approaches. She continued to study metalloproteins using crystallography experiments and molecular dynamic simulations. After MIT, she spent a year as a postdoctoral fellow at Los Alamos National Laboratory.
In 2017, she joined the Hauptman-Woodward Medical Research Institute in Buffalo, New York, as HTX Center Director. There, she also runs a lab continuing work on metalloproteins.
In one project, she has been looking at proteins that help pathogenic bacteria scavenge metals from host cells to wreak their havoc on health. One particular protein they are studying comes from Burkholderia pseudomallei, a bacterial species increasingly found in contaminated soil and water in warming regions of the planet that can cause disease in people and animals.
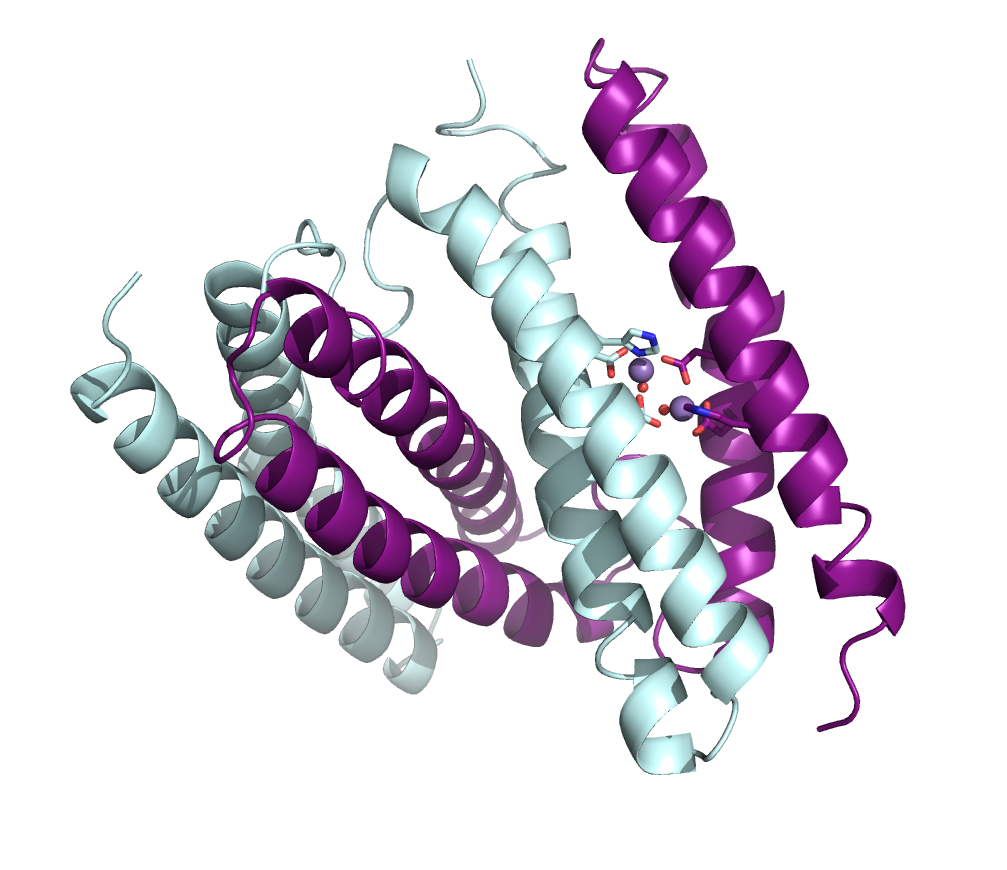
The bacteria can function in both anaerobic and aerobic environments. In its aerobic capacity, it may be reducing peroxide in cells, and Bowman’s team is exploring how that happens.
As a bonus, the protein will form crystals of different sizes, suitable for different types of data gathering. “It turns out to serendipitously be a really great system to work on because we can tune the crystal size,” she says. “So we can get submicron crystals all the way up to 400 micron crystals.” The lab has already solved three structures from X-ray cryocrystallography experiments at the National Synchrotron Light Source II at Brookhaven National Laboratory in New York. They have plans to follow up with experiments to study metal binding, peroxide binding, and dynamics of proteins from B. pseudomallei using room-temperature crystallography, serial synchrotron crystallography and microED techniques.
These days, “structural biology in general is much more integrative,” Bowman says. “It seems like people are much more willing to go, ‘Oh, I really just want to figure out how this works. And so I am going to try this new technique and see if I can get it to work to answer my questions.’”
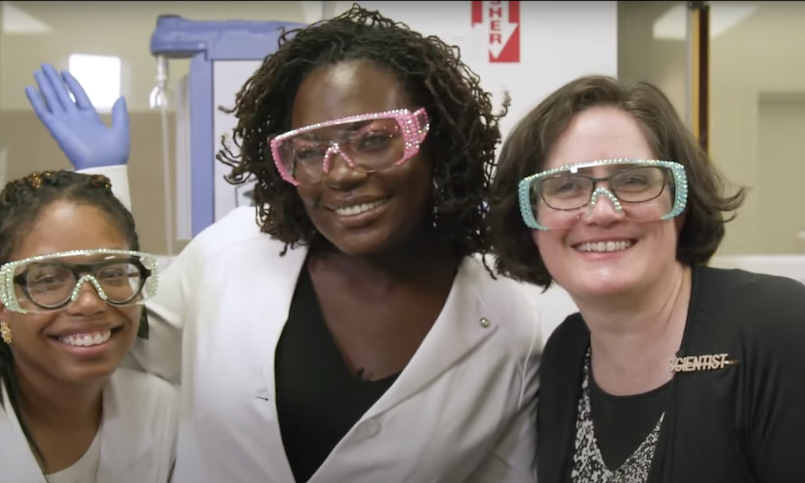

Bowman places a high priority on getting others excited about the science too. She gives talks at community colleges to raise the visibility of structural biology. She has donned crystal studded safety googles with Dr. Raven ‘the Science Maven’ Baxter, a science educator and communicator who runs a popular science vlog. And she encourages students to get in the lab and experience the joy of asking and answering questions.
- Carol Cruzan Morton