Flipping the Switch
Jacqueline Cherfils
École normale supérieure (ENS) Paris-Saclay
Published April 27, 2018
After graduate school, Jacqueline Cherfils attended a Jacques Monod Conference, a prestigious small scientific meeting in France named after a Nobel Laureate. She knew no one. Her goal: To find an interesting molecule to study in atomic detail. She listened to researchers talk about how proteins enter and exit cells. In the end, proteins called small GTPases caught her attention and have captivated it ever since.
“They are famous in biology, because they function as molecular switches in almost every living organism,” says, Cherfils, a National Center for Scientific Research (CNRS) research director at École normale supérieure (ENS) Paris-Saclay.
Small GTPases are so ubiquitous that even thinking about them uses them. To communicate with each other, neurons rely on certain GTPases to fire off stored neurotransmitters. The most well-known GTPase may be RAS, first discovered in the 1980s as an oncogene, a switch that propagates a growth signal through the cell and that can get stuck in the “on” position in mutations that lead to cancer.
At the conference, a well-meaning cell biologist told her the important structural work on GTPases had already been done. After all, this was the early 1990s, and the first atomic structures of RAS had been solved. In fact, structural biologists were just getting started. Since then, more than 500 GTPase structures, alone and in combination with other molecules, have been entered into the Protein Data Bank, including dozens from Cherfils’ lab.
Inside a cell, small GTPases bind a specific chemical—GTP—and from there assemble large molecular complexes that take care of cell logistics. As key regulators of nearly all fundamental cellular processes, GTPases can quickly move between “on” and “off” to make things happen. They may act and look alike, but these versatile proteins have many distinct and specific roles in cells. Not surprisingly, their dysfunction underlies the pathogenesis of many human diseases, such as cancer, cardiovascular disorders, allergies, infections, aging, neurodegeneration, and developmental syndromes.
“We are interested in understanding how GTPases work when cells are fine and normal and healthy, and we’re interested in understanding why and how they are impaired in diseases,” Cherfils says. “From this knowledge, we hope to contribute to drug discoveries by inspiring strategies to inhibit these proteins.”
Cherfils came to structural biology by way of mathematics and computer modeling. She grew up in a small town in the south of France, far from the big research laboratories in Paris she now calls home. Her parents were teachers, a career path she started down before reconsidering her future.

She fell in love with the satisfying logic of mathematics, and pursued it all the way to a masters degree at University of Marseille. “At the level I was studying, everything falls into place in a marvelous, almost artistic, manner,” she says. In college, she noticed a disconnect between equations and the “real world.” And, while she adored her math teachers, most of her friends did not. “I didn’t like the idea of entering a job where most kids are scared of you,” she says.
She jumped to computer sciences, working on mathematics languages and game theory, then moved to Paris to develop tomographic image analysis in a cancer hospital. It was the mid-1980s, the beginning of computer graphics for proteins in biology. She began writing software for early three-dimensional graphic modeling. Then, she became interested in the protein structures themselves and the molecular dynamics of how they interacted in computer simulations. After her PhD in biophysics, she was recruited by CNRS, which funds researchers in France at various institutions and levels, to work on modeling of protein dynamics and interactions.
She spent several months as a visiting scientist in Cambridge, Mass., USA, to work on the molecular dynamics of allostery in a project involving both the Harvard University labs of Nobel Laureate William Lipscomb and Martin Karplus, who would win a Nobel a dozen years later. She made friends with postdoctoral fellows in the neighboring structural biology lab of Stephen Harrison. This played a role in her decision, back in France, to move into a wet lab and generate the structural information herself.When she first identified small GTPases as her research interest, Cherfils interpreted the advice to avoid small GTPases as evidence of a knowledge gap between cell biologists and structural biologists. “I felt like I was from the planet Mars,” says Cherfils, who herself was struggling to understand microscopy studies. When she started her lab, she joined forces with a cell biologist to help make the connections between the molecular details they were uncovering and the more complex processes they triggered.
Cherfils worked her way into studies of the Arf, Rab and Rho families. The Arfs were first discovered as a key protein that cholera hijacks to release the cholera toxin in infected people. Now they are known to control most aspects of intracellular traffic. An early Arf project came from French biophysicist Marc Chabre and his colleague Pierre Chardin. One of Chabre’s papers had inspired her to use aluminum fluoride to image a kinase reaction in a crystal (published in PNAS in 1996). When she read his paper, she had sent him a grateful note. Two years later, he called her with a proposal to crystalize a molecule his lab discovered, which contains a Sec7 domain that promotes fast activation of Arf1. That resulted in a 1998 Nature paper and a welcome mentor and friend for Cherfils.
A milestone in the Cherfils lab came when they succeeded in locking down Sec7 and Arf1 for X-ray crystallography analysis. The paper was important for 2 reasons. First, it showed how a family of GTPases, Arf, is activated in a sequence of events. Second, it showed another way a drug could work to block activity of disease-causing GTPases.
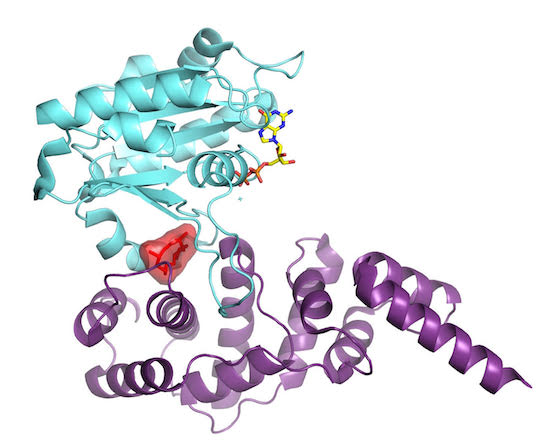
“It turns out that Arf GTPases have a fascinating structural device that allows them to perceive which nucleotide is bound to them,” Cherfils says, “and simultaneously, whether or not it is bound to the membrane.” This system explained how the cell membrane and the activator come together to trip the switch on this GTPase. It was more complex than biologists had imagined. The findings were published December 3, 2003, in Nature.
As they learned how the GTPase protein turned on, they also realized they had found a way to turn it off: Stitch the proteins together ineffectively, as they had done in their experiments. To halt the action at different points for crystallization, Cherfils’ team had used Brefeldin A, a natural compound popular with cell biologists for deciphering the dynamics of the Golgi. It works by stitching the Arf GTPase and its activator together, like double-sided tape. Now stuck, Arf cannot make the further moves necessary to complete activation.
Traditionally, pharmaceuticals are targeted to a specific binding site, blocking the protein-protein interaction by “competitive inhibition. In the case of the Aft GTPase, the proteins were correctly bound together, but they could not move into an active state. “Conceptually,” she says, “we learned from this story that there are other ways protein interactions can be inhibited.”
Cherfils compares it to the martial arts she studied. Drugs are small. Proteins are big. It’s David versus Goliath. “As in martial arts, where one uses the strength of the opponent, the small molecule uses the energy of the protein complex,” she says.
Cherfils’ lab has been working on parallel tracks since then. In the basic science, her team is probing the inner working of these proteins, their activation, and their response to membranes and other proteins.
Over the last few years, biologists have realized that small GTPases should be considered in the environment of the membrane where they work, she says. The next challenge is to understand in molecular and structural detail the ability of small GTPases and their molecular complexes to perceive and probably influence local changes in membranes, she says.
At the same time, Cherfils and her colleagues are interested in the many ways that small molecules can interfere with GTPases. “Nature uses the martial art strategy more than once to block protein-protein interactions,” says Cherfils, referring to plant-derived tubulin inhibitors as an example. “It is exciting that the pharmaceutical industry is now delving into such strategies. We call it ‘interfacial,’ but it is also called ‘stabilizer.’”
Cherfils relishes the entire discovery process—building hypotheses, conducting experiments, and bringing the observations together in some logical, unbiased manner. “There’s a mathematical flavor to it,” she says. She may study protein-protein interactions, but Cherfils has also come to see her laboratory as making connections between people, and the international nature of science as bringing nations together.
Her lab continues to grow crystals and shoot them with X-rays. She has been expanding biophysical and spectroscopic methods and would like to move forward with cryo-electron microscopy to better incorporate membranes into her studies. On the personal side, she has added photography of people, nature, and landscapes to her toolkit, after decades of interpreting atomic protein images.
--Carol Cruzan Morton