In Search of…New Drugs
Doug Daniels
Broad Institute
Published September 30, 2014
When Doug Daniels finished his chemistry degree at the University of Michigan and set off for The Scripps Research Institute for graduate study, he’d already made a key career decision. "I decided I was more interested in the discovery and development of drugs than in the practice of prescribing them," he says.
Having ruled out medicine or even an MD/PhD program as a way forward, he instead dove into structural biology. He’d come to love organic chemistry in college, but it was the visual aspects of how structure relates to reactivity that hooked him. "I wanted to take that same kind of thinking and apply it in a larger structural space, at the protein level," he says.
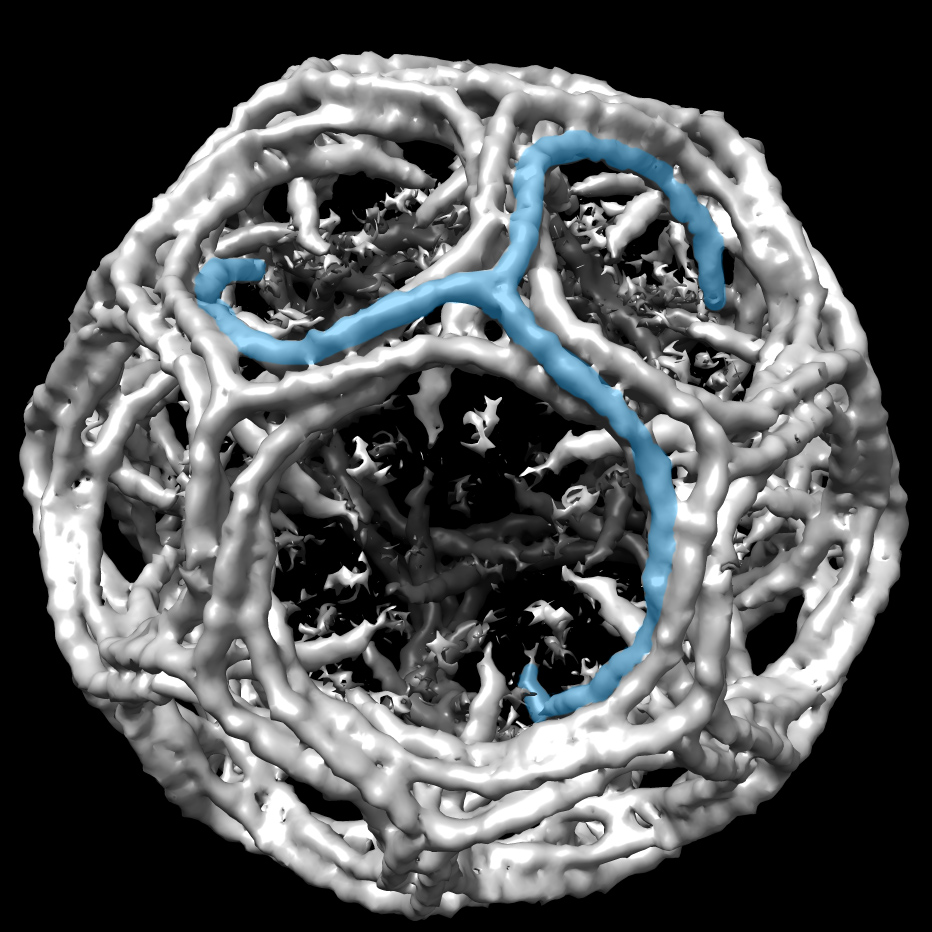
The transition led him to study DNA repair proteins. The project, done in collaboration with a medical researcher, delved into O(6)-alkylguanine-DNA alkyltransferase (AGT), a DNA repair protein that is hijacked by cancer cells as a way to resist DNA-damaging chemotherapy agents. Working in the laboratory of John Tainer, Daniels solved the structure of AGT alone and in complex with two inhibitors that were in early development as drug candidates for combating resistance.
Daniels followed this training with a post-doc at Yale University. Instead of continuing to pursue structural biology and proteins, he switched gears again. "I started to become interested in the peptide as the drug," says Daniels, who joined the laboratory of Alanna Schephartz, who was working on peptide engineering.
Daniels and others in the lab worked on beta-amino acid peptides. These peptides are formed with amino acids that differ from naturally-occurring amino acids in that they have an extra backbone methylene unit. "Because of the longer backbone, they have completely different structural characteristics," says Daniels. For instance, they join together to create helices, as natural amino acids do, but not the same kind of helix.
As if playing with a new kind of Lego set, Daniels and colleagues assembled structures using beta-amino acids, first building helices, then forming an octomer of eight helices. Using his crystallography background, Daniels solved the resulting structure and found that the helices had associated to form a globular protein-like structure, complete with protein-like folds. Having achieved this, says Daniels, “you can start to think about it as a starting point for the design of peptides with specific enzymatic functions,” something Schephartz has made progress on recently.
Daniels' skill set attracted the notice of Novartis, and in 2008 he joined a group in the Cardiovascular and Metabolic Disease department working on protein design for protein therapeutics. For instance, he worked on a protein therapeutic for diabetes, using protein engineering to enhance the activity of the protein.
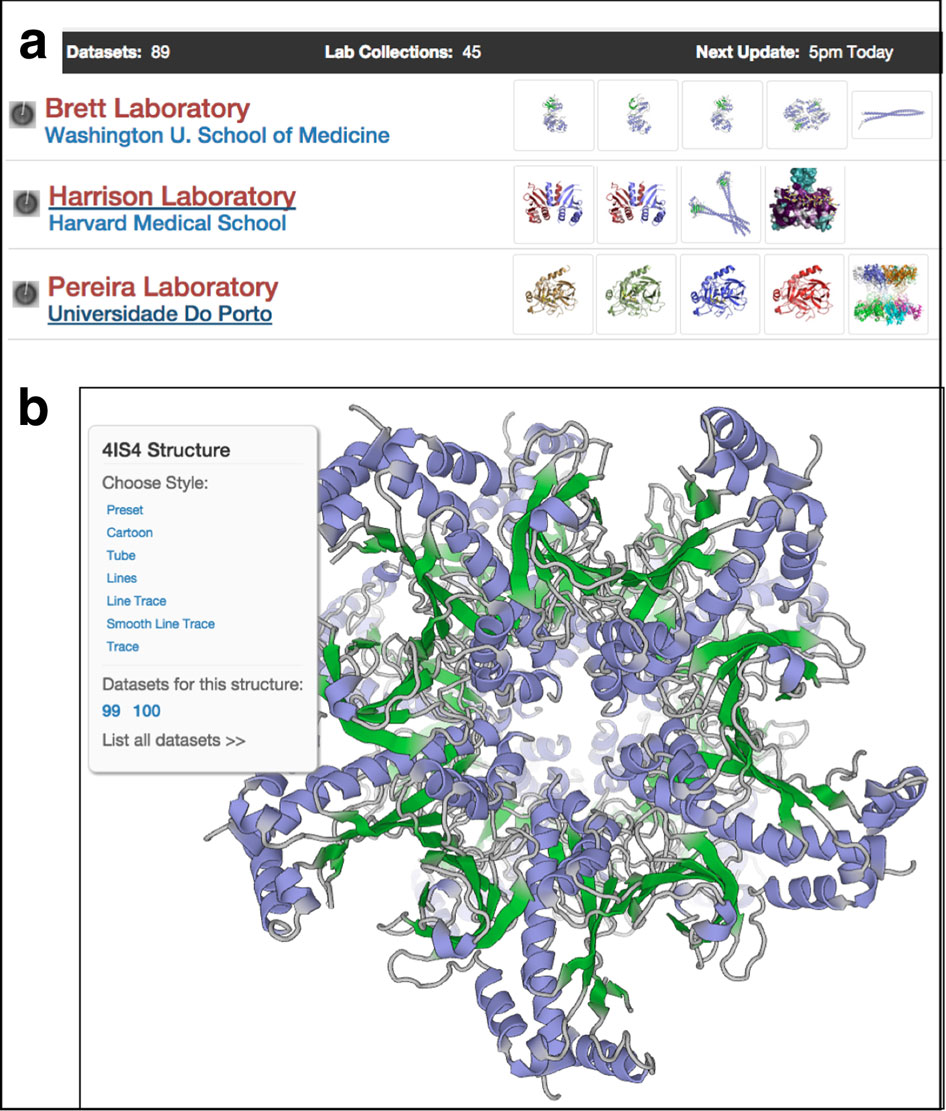
In 2012, Daniels moved to the Broad Institute, which was applying genome-inspired science to drug discovery and development. Daniels’ first task was to build a protein science group, which later expanded to include structural biology. "Having structural information in the drug discovery process is priceless," he says. "We use compound-bound structures to rationalize and design improvements in our compounds."
Daniels has his protein sciences team assembled and in just one year got his crystallography group running full speed. "SBGrid helped us get up and running with the software with very little down time," says Daniels.
The team is focused on discovering new drugs for targets thought to be "undruggable," such as PCSK9 for coronary artery disease, and MCL1 and ERG for cancer. "If we thought these targets were undruggable, we wouldn’t be pursuing them," says Daniels, who expects to be publishing his team’s first set of findings soon.
-- Elizabeth Dougherty