Charm and Diplomacy
Gerard Kleywegt
EMBL-European Bioinformatics Institute, UK
Published March 7, 2012
“I was an angry young man,” says Gerard Kleywegt of his early days in the 1990s as a structural biologist. He’d found his way from the University of Utrecht, in the Netherlands, where he’d done his PhD on Nuclear Magnetic Resonance (NMR) spectroscopy, to Uppsala, in Sweden, where as a young post-doc he was learning X-ray crystallography from Alwyn Jones. “I thought quality and validation of structures was so important that, when I found an error, I was almost shocked.” And he wasn’t quiet about it.
Kleywegt now laughs at his “zeal,” and says, “I’ve mellowed a lot.” But the nickname Jones gave him, “CD,” has stuck. It stands for “Charm and Diplomacy” — the two least likely characteristics to be associated with Kleywegt, according to Jones.
Kleywegt’s passion for validation has also stuck. Today, working at the EMBL-European Bioinformatics Institute (EBI) as Head of the Protein Data Bank in Europe (PDBe), he oversees a major effort to improve validation throughout the PDB, a role that every turn of his career has prepared him for.
Originally, when Kleywegt went to Uppsala, he expected to spend just a year or two there learning crystallography before going on to a career in drug design. “I ended up staying for nearly 18 years,” he laughs. “I really enjoyed it. It was a very dynamic lab.”
While in Uppsala, he solved structures and wrote software tools, mainly to meet his own crystallography needs. He created what is now known as the Uppsala Software Factory, also tongue-in-cheek, he says, “because the so-called factory was just me.”
But what Kleywegt is probably best known for are his efforts to improve and advance structure validation. In fact, when he took on his new role at PDBe, he was already a member of the Worldwide Protein Data Bank (wwPDB) validation task force for crystallography. “When I came here, I transitioned from being an advisor to advisee,” he says.
The goal of the wwPDB validation effort is to make it easier for scientists who are not experts in structural biology to select structures intelligently from the PDB. This requires automating a number of very complex processes. To do this, task forces in each field (i.e. NMR, X-ray crystallography, EM) recommend validation methods and tools, which the wwPDB then knits into a “validation pipeline,” explains Kleywegt.
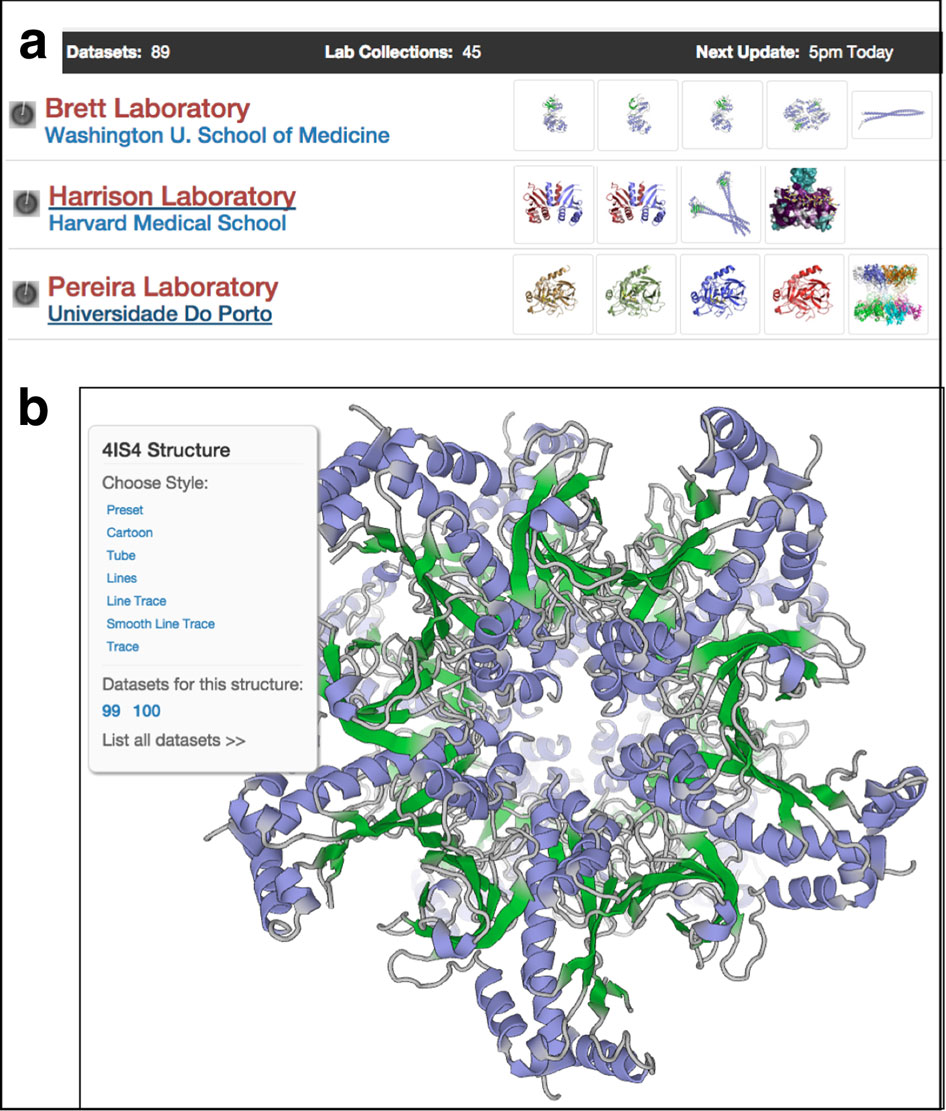
This automated pipeline consolidates results and assigns each structure a score to indicate its relative quality. Scores appear on a simple, intuitive sliding color scale from red (poorer scores than other structures) to blue (better scores). Each newly deposited structure gets validated and scored in the same way. Depositors are informed of possible errors or concerns that may need closer scrutiny, and are given an opportunity to address the issues before the data are made public. “Hopefully, this will prevent errors from making it into the archive in the first place,” says Kleywegt.
Though Kleywegt no longer has time to write software, which used to be his reward for completing other work, he says (with both charm and diplomacy): “This job is ideal for me, with my background in chemistry, NMR, crystallography and bioinformatics. But much as I love working here, I miss Sweden every single day.”
-Elizabeth Dougherty