New Kid on the Block
James Chen
Oregon Health and Science University
Published July 29, 2014
James Chen was raised by mathematicians who taught him at an early age to program computers and to think analytically. “Everything had to be formulated. Instead of speaking in natural language, we sometimes spoke in formulae at home,” Chen says with a laugh. No surprise, then, that Chen, assistant professor of biochemistry and molecular biology at the Oregon Health and Science University (OHSU), became an expert in electron microscopy data analysis.
Physics appealed to him as a college student in China, but as a graduate student at Florida State University, he found himself lured into biophysics. He began working with Michael Chapman, now also at OHSU, on an X-ray crystallography project to perform structure refinement in real space — against the electron density map — rather than in reciprocal space.
Instead of going straight to a laboratory as a post-doc, Chen followed his PhD with two years of computer science training, earning a Master’s at the University of North Carolina at Chapel Hill.
Then, in 2003, he began a post-doc at Brandeis University in the lab of Niko Grigorieff focused on methods development for electron cryo-microscopy. Chen developed a program called SIGNATURE, which automatically screens for particles, speeding the process of particle selection and increasing data set sizes by ten-fold or more. “Single particle data analysis is all about statistics,” says Chen. “The larger the dataset the better.”
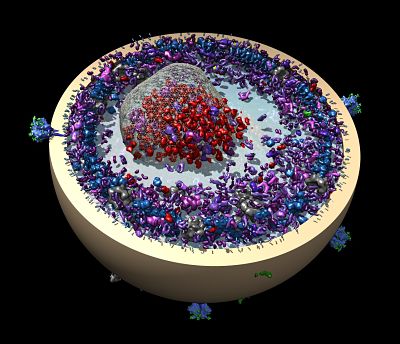
He also worked on a project in collaboration with Stephen Harrison at Harvard Medical School to solve the capsid structure of the rotavirus, reaching 4 Ångstrom resolution, one of the highest resolution EM structures at the time.
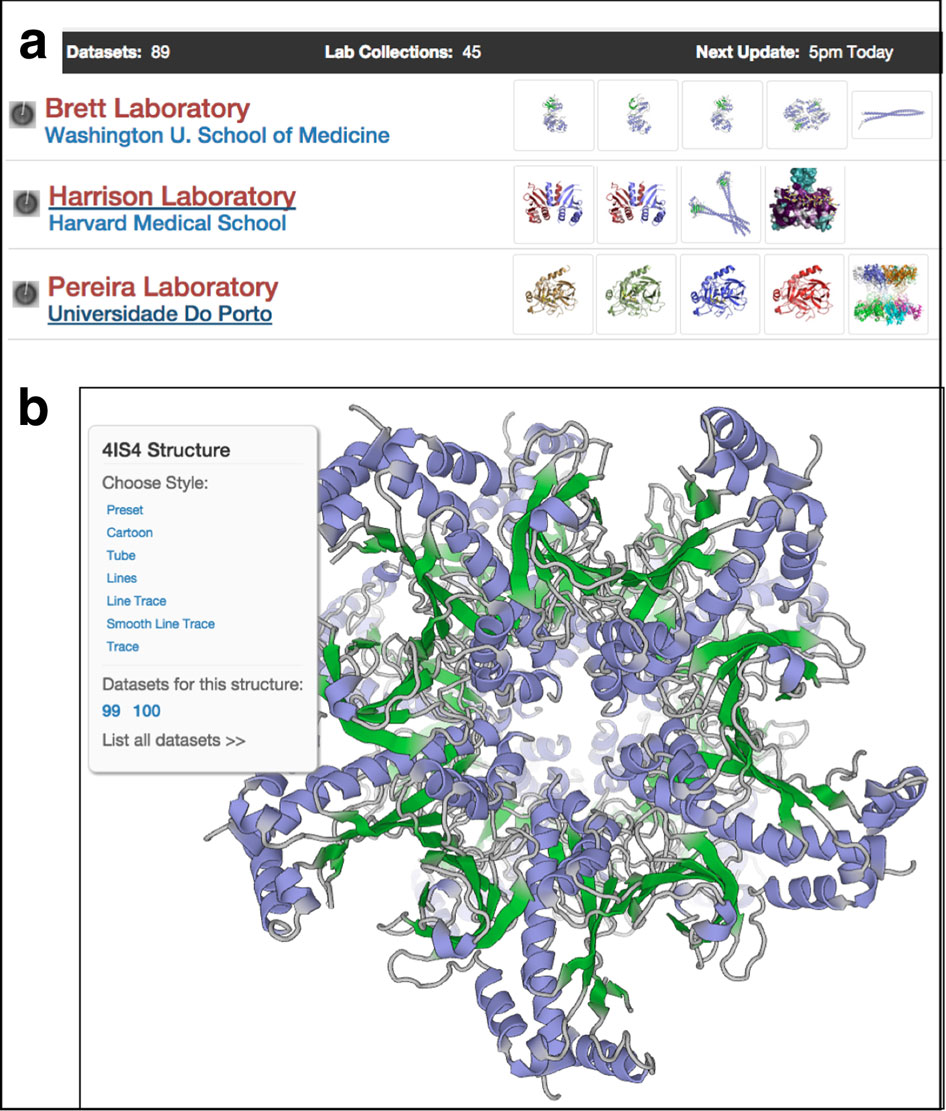
Chen moved to MIT in 2011 where, as a research scientist, he set himself with the task of taking a fresh look at single particle EM data analysis, down to its theoretical foundations. His goal was to put together a comprehensive and user-friendly computing environment for end-to-end EM data analysis to make it easier for researchers to test their ideas, he says, “at the speed of thought.”
The end result of his efforts is PARTICLE, which has been used by a few labs including his own. “It’s really the new kid on the block," says Chen, who hopes to be able to publish a high-resolution structure based on the platform in the near future.
In November 2013, Chen moved to OHSU to start his own lab. He has three complementary research focuses. The first involves technical efforts to advance EM data collection, with focuses on low dose-rate imaging techniques and fast data collection methods, including high-throughput tilt-pair data collections. The second is methodological work to improve EM resolution, with a focus on breaking the physical barriers to achieve super-resolution EM.
Chen is fortunate to have support from two neighbors, nearby microscope manufacturer FEI, which sponsors and supplies equipment to the OHSU Living Lab, and Intel , which is providing him with access to supercomputing resources.
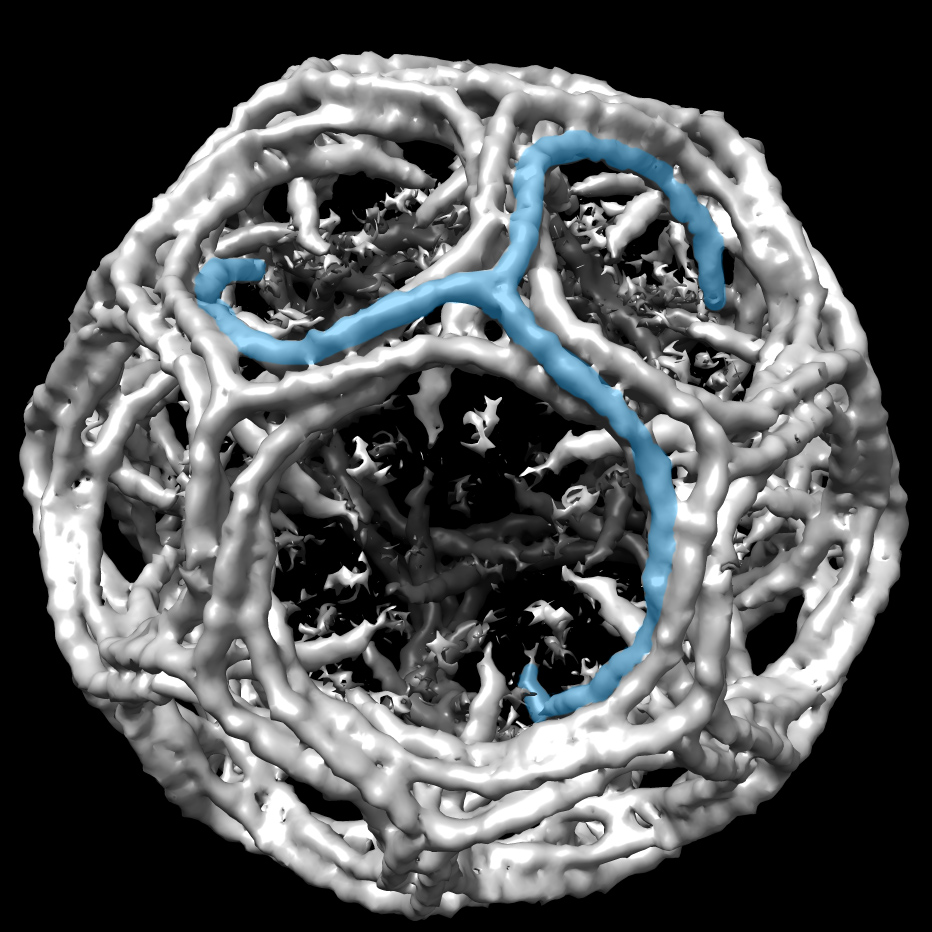
Finally, Chen is applying his new methods to structural biology, with efforts underway to understand the structural detail of the nuclear pore complex, the gatekeeper between the cytoplasm and the cell’s nucleus. “We do technical and methods research, but the proof of concept has to come from the application,” he says. “The three parts are tightly connected. Improvement in one area will certainly lead to development in the others.”
Chen has hired a post-doc and a lab manager, both already hard at work on his projects, and he hopes to attract more post-docs to his lab. “This is the golden age of EM,” he says. “With the resources we have here in Portland, this is a very good place to develop a career.”
-- Elizabeth Dougherty