Turning the DIALS
Nicholas Sauter
Lawrence Berkeley National Laboratory
Published June 29, 2016
Nicholas Sauter began working on DIALS (Diffraction Integration for Advanced Light Sources) in 2011 because he and his colleagues recognized that the experimental methods of X-ray crystallography were changing, and changing fast. To be usable, the software that automates crystallography experiments must be able to keep up.
So he and his team at Lawrence Berkeley National Laboratory and collaborating teams at CCP4 and at the Diamond Light Source synchrotron in the United Kingdom developed a modular system that allows new algorithms to be dropped in as new experimental methods and technologies emerge. Examples include handling data from faster detectors, like the Pilatus, handling new technologies, such as the X-ray free electron laser (XFEL), and handling new types of experiments, such as putting multiple crystals in the beamline at the same time, or running experiments using two different wavelengths at the same time.
Since DIALS is both modular and open source, new units can be added at will. “There are many things you can do that are very flexible if you’re able to write a little bit of code,” says Sauter, Computational Staff Scientist at LBL. “Or you can talk to us and we can write the code for you.”
Sauter first discovered the allure of structural biology as a high school student in New York City. In a tenth grade honors biology course in 1978 he learned about the structure of myoglobin, which had been recently solved. Then in college at the University of Chicago, he did a lab rotation in the laboratory of pioneering structural biologist Paul Sigler.
As a graduate student at Harvard University in the late-1980s, Sauter joined the lab of another famed structural biologist, Don Wiley. In the Wiley lab he worked to apply nuclear magnetic resonance imaging to measure a binding constant the team was interested in. “Wiley gave me the space to become an expert on something that was new to the group,” says Sauter. “At the time, crystallographers did crystallography, but there was an appreciation that we needed to bring in new methods.”
Sauter did a post-doc at the University of San Francisco working on glucocorticoid receptors, but his focus quickly shifted to computation. In the 1990s, Sauter moved to the Stanford Linear Accelerator Center where he took a position as a collaboratory scientist at the Stanford Synchrotron Radiation Lightsource. At that time, the crystallography field was becoming increasingly automated, to the point that the experiments themselves becoming remotely controllable. “Things were coming together so that the whole protein crystallography experiment could be controlled by a piece of software sitting at a single terminal,” says Sauter. “It could all be done through software.”
The Stanford team developed BluIce, a user interface to facilitate such remotely controlled experiments. Sauter added code to show the diffraction patterns to the remote user as the data were acquired, and to illustrate the experimental parameters under user control. The BluIce beamline control system is still in use today.
In 2000, Sauter moved to LBL to work on what would become Phenix. But because of his beamline experience, he was pulled into work focused on high-throughput crystallography, and, naturally, more automation. As part of this work, he contributed LABELIT and Spotfinder.
About a decade later, light sources were becoming brighter, detectors were becoming faster, and experiments were becoming more complex and data intensive. Sauter and colleagues recognized the need for new software to interface with these new experimental tools and methods.
With funding from the National Institutes of Health at LBL, and from the the European Union’s BioStruct-X and the Wellcome Trust in the UK, Sauter and colleagues began a four-year effort to develop DIALS. Their first step was to develop a modular system using Python and C++ that would allow modules to be run as needed. They then implemented existing methods, familiar from programs like d*Trek, Mosflm, XDS, and EVAL15, as drop-ins to DIALS. The software is distributed with CCP4 and also available with XIA2.
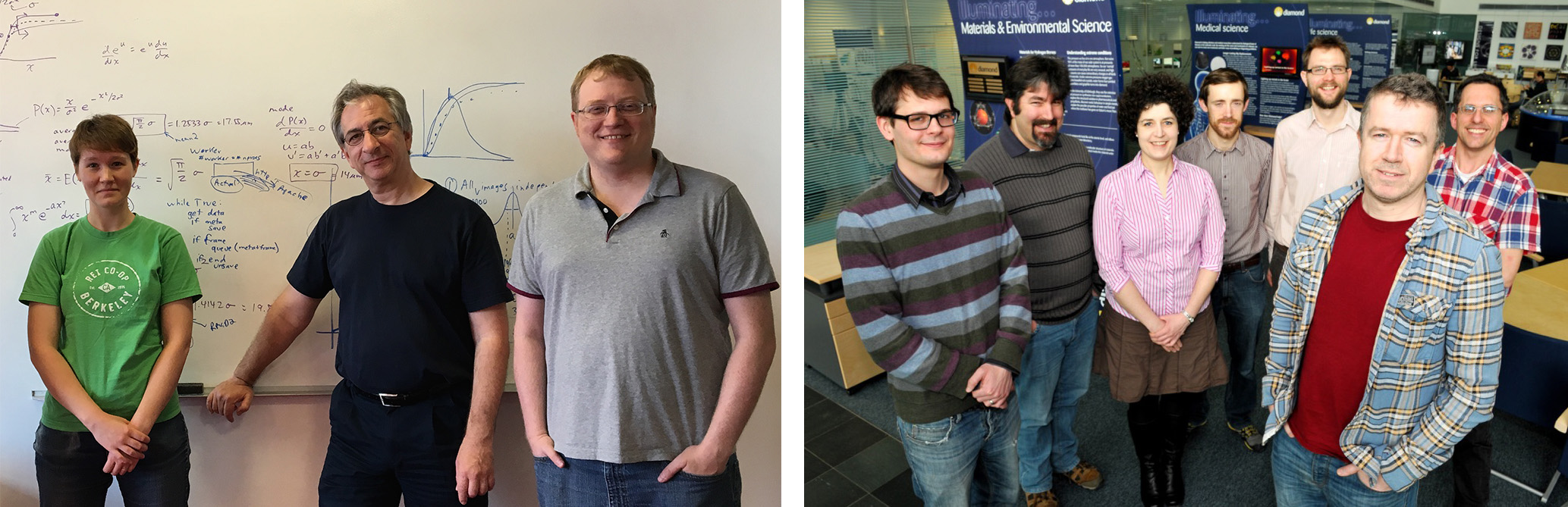

Left: the LBNL developer team: Iris Young, Nick Sauter, and Aaron Brewster.
Right: Diamond Light Source developers: James Pankhurst, Luis FuentesMontero,
Melanie Vollmar, Richard Gildea, David Waterman (from CCP4), Gwyndaf Evans, and Graeme Winter. Not shown: Marcus Gerstel.
The group is currently applying DIALS to support XFEL experiments. These use femtosecond X-ray pulses to avoid radiation damage to samples, and the sample is examined at room temperature. “XFELs allow you to see protein conformations and fluctuations under physiological conditions,” says Sauter. “So this gives you a framework for understanding protein regulation at room temperature.”
The team in collaboration with David Eisenberg at UCLA, is also using XFELs to look at crystals produced inside live cells, including a protein toxin that is produced as a crystal inside a bacterium. “New technologies open up interesting possibilities,” says Sauter. “It’s just another way to do crystallography that will allow us to do things that are important.”
-- Elizabeth Dougherty