
Second Takes
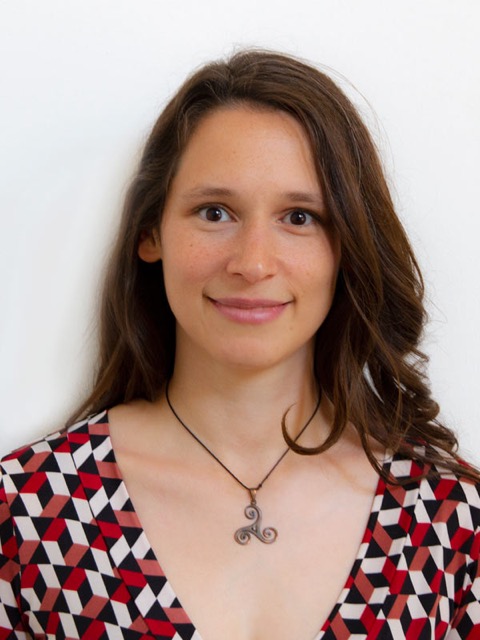
Andrea Thorn
University of Hamburg
Published February 28, 2023
Andrea Thorn had been a junior group leader for a less than a year when, in early January 2020, researchers in China identified the cause of a mysterious contagious illness with pneumonia-like symptoms. Less than a week later, they released its genetic code, which revealed the novel virus to be a close cousin to the SARS coronavirus (SARS-CoV-1) that caused an outbreak in 2002-2003.
The rapid scientific response gave Thorn an idea for a talk she was due to give over coffee and cake to colleagues in her building at the Julius Maximilian University of Würzburg …
Find out More »













 Bob Langridge. Photo by Christopher Springmann, …
Bob Langridge. Photo by Christopher Springmann, …













