Particle catcher
Stefan Raunser
Max Planck Institute of Molecular Physiology in Dortmund, Germany
Published June 28, 2021
If insects had horror movies, one of the scariest would feature tiny monsters. The grisly drama would spotlight the real-life threat to insects from young threadworms, smaller than eyelashes, and their symbiotic gut bacteria. The worms seek out and penetrate insect larvae. Inside the larvae, the worms release their gut bacteria, Photorhabdus luminescens, named for their mysterious glow. The bacteria take over, emitting a slew of toxic proteins that quickly kill the insect. The bacteria thrive on the insect cadaver. In turn, the worms feast on the multiplying bacteria, reproduce, and move on to the next larvae.
The setting would be a farm, because the worm and its bacteria are deployed as biological pest controls. But one of the bacteria’s toxic complexes has potential for medical use in people, according to Stefan Raunser and his team at the Max Planck Institute of Molecular Physiology in Dortmund, Germany.
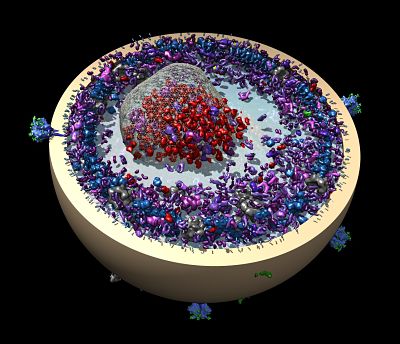
Raunser’s team has worked out key molecular details for one of the bacteria toxins. A toxin complex (Tc) in the form of tiny molecular syringes pierces cell membranes to inject a deadly toxin. (The same toxin complex is found in the plague bacterium Yersinia pestis, but it’s not clear if it is involved in the pathogenicity.)
There are more questions about the toxin complex (Tc), but Raunser realized the spring-loaded molecular injection complex may have potential as a biological medical device.
His team reported a step toward repurposing the lethal mechanism to deliver therapeutic proteins (Nature Communications, 2019). They found a way to replace the toxin in the bacteria's nano-syringe with a different protein and inject it into cells. Further research will determine its value in fighting disease, such as cancer.
The toxin complex and its potential drug delivery application was deciphered using the Raunser lab’s signature combination of single-particle cryo electron microscopy (cryo-EM), biochemistry, and original software.
In fact, new biological knowledge is not the only lab product. Since 2014, when he became head of the structural biology department, Raunser has deployed a computer science group to make it easier and faster to do the science.
“There are software developers, and then there are biochemists who want to click and get a result,” Raunser says. “In my lab, both work closely together.”
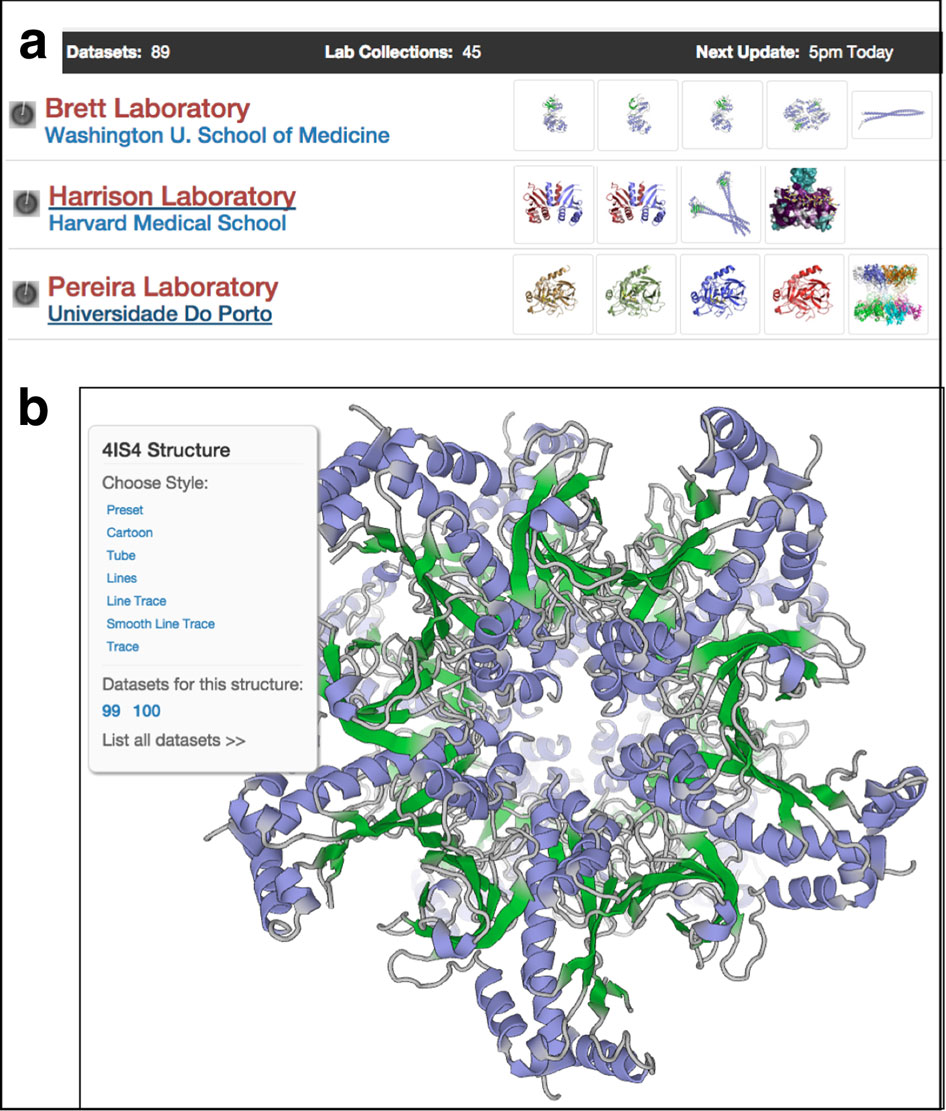
The computer science story goes back to a postdoctoral collaboration with Pawel Penczek, a pioneer in cyro-EM image processing, who developed SPARX. Working on new biology problems in his own lab, Raunser wanted to give the imaging processing software tool a facelift to improve the workflow and accessibility. After all, cryo-EM was becoming a more popular and important tool. They named the new version SPHIRE (JOVE, 2017).
When Raunser launched his in-house developer group, they added an artificial intelligence function to select particles in a package called SPHIRE-crYOLO.
In single-particle cryo-EM, scientists need to select more than 100,000 particles of a molecular complex to put together a detailed near-atomic structure. Then the particles need to be digitally aligned, classified, and reconstructed from the multiple view angles scanned with the microscope.
“In the past we hired a student to pick particles manually in the afternoon after lectures,” Raunser says. “You always work with imperfect samples. It can become very complicated.”
In SPHIRE-crYOLO, postdoctoral fellow Thorsten Wagner delegated the painstaking particle selection to a computer (Communications Biology, 2019). Compared to hand picking, the software was able expertly to distinguish more true particles from icy crystals in the frozen sample grid, resulting in slightly improved resolution in the final structure.
The next year, amid the coronavirus pandemic, the group released TranSPHIRE, led by postdoctoral fellow Markus Stabrin. The automated image processing software acts like a quality-control manager during the image gathering stage. It calculates two-dimensional classes and generates 3D reconstructions during data collection (as fast as 24 hours). The software incorporates a data-picking feedback loop with classification and deep learning (Nature Communications, 2020).
“It saves a lot of time,” Raunser says. In the past, a researcher spent 2-to-3 days collecting data at the microscope, then another few days processing the data to learn what they collected. If the data was not good, it was back to the microscope for another time-consuming round of data collection and evaluation.
“Now with this in hand, you can monitor and collect good data,” Raunser says. “If it’s not good, you can stop and collect at a different region of the grid.”
Raunser grew up in Germany, son of a craftsman (father) and medical assistant (mother). He was the first in his family to earn a high school degree, or abitur, as it is known in Germany. As a boy, he remembers being eager to learn new things to understand nature.
He was also a budding musician who mastered several instruments. Beginning in second grade, he learned to play the accordion. Eventually, he specialized in playing the tango at dances and in restaurants. (Now, he just plays for family.)
Meanwhile, by fifth grade, Raunser knew biology was his future. In college, he decided to become a biology and chemistry teacher.
“It turned out I was more interested in research,” he says. At University of Mainz, he studied plant biochemistry and worked on one of the most abundant membrane proteins in the world, a chlorophyl complex that harvests light for plants. On that project, he met and worked with his future PhD advisor, Werner Kühlbrandt.
In those studies, he discovered how much the three-dimensional (3D) structure of a protein reveals and learned to adore the structural work. For his doctoral work at Max Planck Institute of Biophysics, Raunser dedicated himself to 2D electron crystallography, an early and arduous technique in electron microscopy to study glutamate transporters. He noticed many of his lab mates were switching to X-ray crystallography for the higher resolution it promised. But Raunser felt electron microscopy had a bright future, so he stuck with it.
For his postdoctoral work, he moved to the Harvard Medical School lab of Tom Walz in Boston. He used extended single particle electron cryo microscopy to work on the membrane urea transporter and many large protein complexes.
Raunser had left his accordion in Germany, but during his three years in Boston, he played violin in the Cambridge Symphony Orchestra.
Near the end of his postdoc, electron microscopy was starting to live up to its promise. The resolution was steadily improving, thanks to advances in technology and techniques. However, it was still 5 years before the resolution revolution in cryo-EM.
Then Raunser won a fellowship to establish his own group in Germany in 2008. He continued with single-particle cryo-EM but started work on a completely new project looking at membrane proteins involved in cholesterol homeostasis.

It was challenging and slow. So Raunser was ready to listen when a colleague approached him for a collaboration on a bacteria toxin (Tc) that needed to breach the membrane to enter a cell.
“The system turned out to be extremely interesting,” Raunser says. “That’s how it is in science. You might have a great idea in one direction, and something else comes along that is very attractive, and you pick it up.”
Soon, another colleague asked him to look at another set of molecules, the filament-forming proteins that make muscles contract. Using cryo-EM at first, they captured detailed structures of the key proteins involved in muscle contraction using cryo-EM (Cell, 2012; Nature, 2015; and Nature, 2016). They solved the first high-resolution structure of the ryanodine receptor, a calcium channel on the surface of muscle cells that triggers them to contract (Nature, 2015).
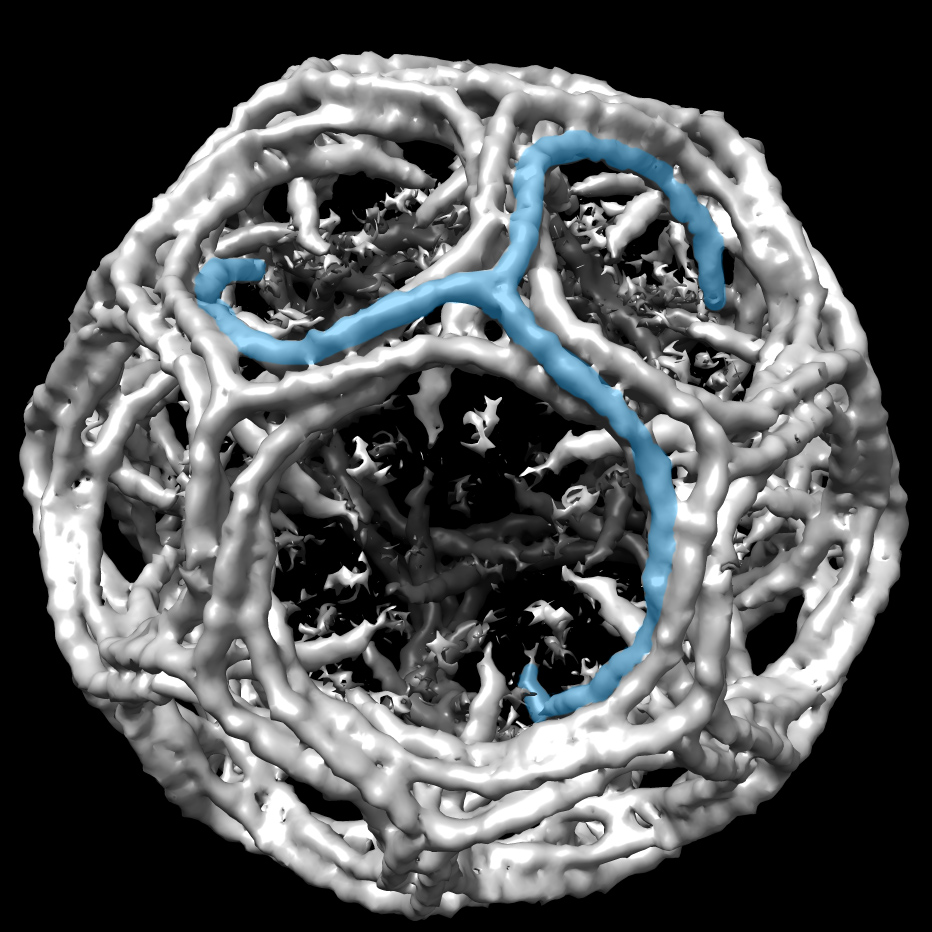
More recently, Raunser’s group has turned toward cryo-electron tomography (cryo-ET), which shows not only proteins but also their spatial arrangement. In a mouse psoas muscle, they showed for first time the 3D organization of the smallest subunit of muscle, the sarcomere (Cell, 2021). They discovered new details about how myosin and actin are organized to give the sarcomere the elasticity to win races and set world records, in the case of elite athletes.
As he learns more about the biology of muscle protein complexes, Raunser plans to investigate what goes wrong in muscle diseases. He’s also curious about what happens in muscle injuries in soccer players and the high-performing muscles of racehorses.
The experimental results with the sarcomere also reflect a new direction of software and hardware development in the lab. Those efforts are now focused almost exclusively on cryo-ET, where Raunser sees more of a need for new tools and techniques to select particles or substructures and improve the workflow of data collection and image processing. The group is also collaborating with companies on the hardware.
“My philosophy is for hardware and software development to come together to solve important biological questions,” Raunser says. “Biology is in the forefront. We want to be able to answer questions.”
-Carol Cruzan Morton