X-PLORer
Axel Brunger
Stanford University
Published October 1, 2012
Axel Brunger joined SBGrid in the early days, in 2006, but he may be best known among structural biologists as the man behind CNS (the Crystallography & NMR System), which he contributes to SBGrid, among other tools. Today, however, Brunger focuses almost all of his work on understanding the molecular mechanism that causes neurons to release neurotransmitters and propagate nerve signals.
"Most drugs for treating neurological diseases affect postsynaptic signaling," says Brunger. "If we could develop compounds that could act on the actual release, that could open up a range of more specific and finely tuned drugs."
Long before exploring neuronal proteins, Brunger made his mark in computational biophysics. In 1987, he released a program called X-PLOR for the derivation of three-dimensional structures using NMR data. This program became the basis for crystallographic refinement by simulated annealing by implementing restraints for X-ray diffraction data.
Later, he and his collaborators developed CNS, a "second generation" system. "We wanted to expand the scope to include all aspects of crystallographic phasing and to create a self-contained system," says Brunger. CNS includes Brunger's refinement methods, such as cross-validation and simulated annealing, as well as density modification methods and methods to obtain phase information by MAD phasing or molecular replacement.
In addition to speeding structure determination and improving validation of crystal structures, CNS also ushered in a new approach to crystallographic programming. "Instead of writing one opaque program that the user can't see into, we decided to develop a scripting language," says Brunger. He and his colleagues at Yale University, where he developed most of CNS, coded their algorithms and methods in scripts and used a FORTRAN program to do the heavy computational lifting.
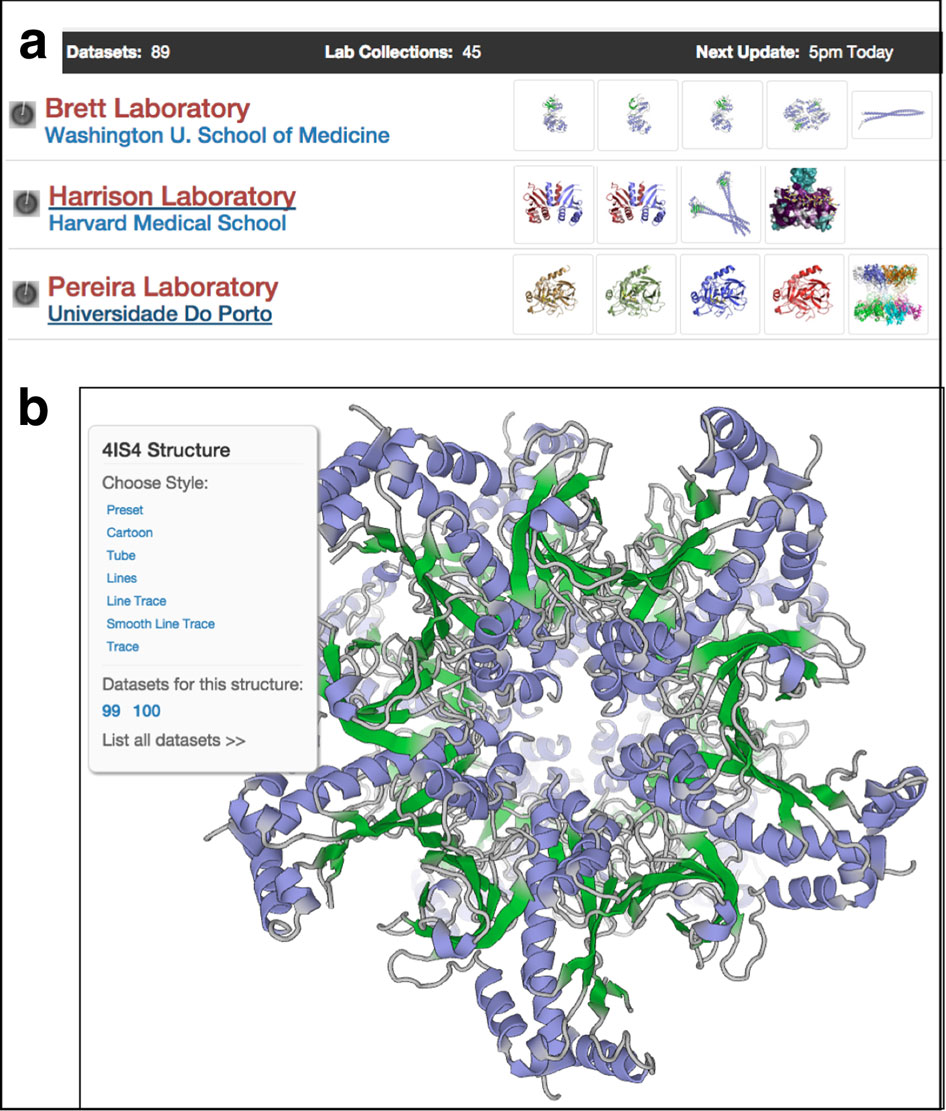
Most recently, Brunger, now at Stanford University, together with Gunnar Schröder, now at the Forschungszentrum Jülich, and Michael Levitt at Stanford, developed a new refinement method, called deformable elastic network (DEN) refinement, and implemented it in CNS. In collaboration with Piotr Sliz at Harvard Medical School, this computationally intensive DEN-refinement method has been made available through the SBGrid Science Portal. "You prepare and test the refinement script file locally, then slip it onto the server. It will submit as many jobs as needed to get the best possible solutions," says Brunger, who with Sliz describes the work in January 2012 issue of Biological Crystallography.
In parallel, in the mid-1990s, Brunger decided to pursue experimental science. "At that time, people had just discovered some of the key proteins involved in neurotransmission, such as the SNARE family of proteins," he says. "But there were no structures for any of them." Having done some work on membrane protein structure prediction, it seemed like a good system to explore. In the late 1990s, he and Reinhard Jahn, now at the Max Planck Institute of Biophysical Chemistry, determined the structures of some of the key proteins and complexes involved in the process of synaptic vesicle fusion.
In 2000, Brunger moved his lab to Stanford University because of its close proximity to the SLAC National Accelerator Laboratory and the establishment of an interdisciplinary initiative called Bio-X. His many recent experimental accomplishments include determining the first structure of a botulinum neurotoxin in complex with a SNARE protein and the elucidation of how botulinum neurotoxin recognizes its target on the neuron.
Brunger also employs single-molecule fluorescence microscopy methods to complement his structural studies. "Crystallography only gets you snapshots," he says. "So we use optical microscopy to get at the dynamics and the molecular mechanism."
While today, almost 100 percent of Brunger's lab focuses on experimental biology, Brunger expects a resurrection of computational work in his lab inspired by the first hard X-ray free-electron laser at SLAC. "It will allow us to look at very small crystals or those that are extremely radiation sensitive," he says. "But there are a lot of challenges around how to optimally analyze and interpret such data."