Pushing the Boundaries
Stephen Harrison
Harvard Medical School
Published April 22, 2013
In the mid-1960s when Stephen Harrison began to determine the structure of the tomato bushy stunt virus, SBGrid didn't exist. There was no need for it. They didn't even have a hard disk for storage.
"Near the end of the 60s they got a disk. That was a big deal," recalls Harrison, Giovanni Armenise-Harvard Professor of Basic Medical Sciences at Harvard Medical School. "One disk."
Lacking storage and a network, as a doctoral student in biophysics at Harvard, Harrison had to walk to the Computer Center to code his programs on punch cards. The limiting factor, however, wasn't so much the burden of manual labor as the cost of CPU time. The billing algorithms for the mainframes scaled with a program's use of random access memory (RAM), which by the early '70's still maxed out at a megabyte.
"There was a big premium on writing programs very efficiently and over-writing locations the instant you no longer needed the information," says Harrison, who worked with punch cards and mainframe technology through to 1977, when he completed the final computations of the structures of the bushy stunt virus at better than 3 angstroms resolution.
"One could not have done it more than a year or two earlier," says Harrison. "We pushed forward more or less as fast as the computing resources allowed."
Laboratory-based DEC and VAX computers debuted in the 70s, accelerating the pace of science. "We could do our day-to-day computations in the laboratory, and we could do them over and over again and make mistakes without pouring money down the drain," says Harrison.
But the DEC computers that ran film scanners were still so RAM constrained that they could not hold a scan of the film used to collect the X-ray data, which, at the time were recorded using conventional laboratory X-ray generators. Harrison's programs did piecemeal scans, doubling back when necessary.
Later, in the 1980s and into the 1990s, it became feasible to record crystallographic data at shared synchrotrons, and X-ray detection transitioned from film to electronics and, more recently, direct pixel array detectors, such as the PILATUS collector used at the Northeastern Collaborative Access Team (NE-CAT) facility at APS. NE-CAT is a shared synchrotron dedicated to X-ray crystallography at the Argonne National Laboratory, managed by Cornell University, and shared by researchers, including Harrison, at multiple institutions in the northeastern US.
Today, says Harrison, biological problems in X-ray crystallography are, for the most part, no longer computationally limited. As a result of the recombinant DNA revolution, neither are they biologically limited by the need to find a purifiable protein or complex of interest in large quantities in nature.
Rather, says Harrison, "they are limited by whether or not we can find an embodiment of an interesting biological problem in a molecule that crystallizes."
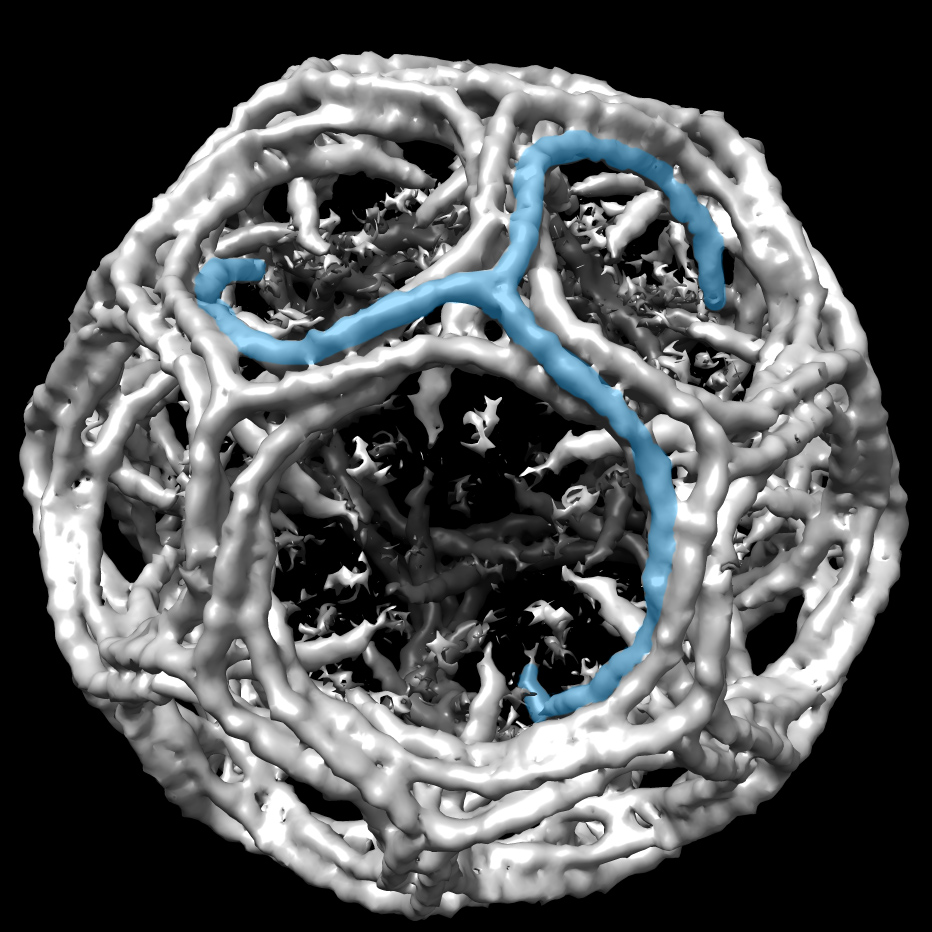
As a result, some of Harrison's current projects, related to human virus structures, are moving away from X-ray crystallography toward electron cryomicroscopy (EM). EM technologies are advancing rapidly and, with the recent introduction of direct electron detectors, reaching a new level of maturity. "I think over the next five years the boundary will move toward electrons and away from X-rays for problems that have no electron solution today," says Harrison.
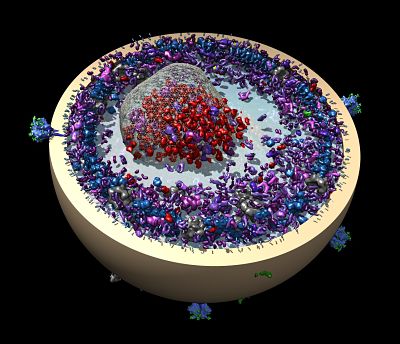
As an example, Harrison is trying to use EM to investigate interactions of human antibodies with the influenza hemagglutinin glycoprotein. As an immune response develops, antibodies mutate toward higher affinity. Understanding such antibody affinity maturation is a key part of vaccine development.
If EM proves to be a more straightforward and rapid way to solve these structures, Harrison speculates that EM may make its way into the vaccine development process much the same way X-ray crystallography has become an essential tool for drug development.
Meanwhile, EM image analysis is pushing the boundaries of available computing, says Harrison. He is, once again, moving forward as fast as the computing resources allow.
-- Elizabeth Dougherty