Shape-shifting secrets of membranes
James Hurley
University of California, Berkeley
Published November 27, 2020
The story of how a viral protein meets a cell membrane has become a compelling narrative during the COVID-19 pandemic. James Hurley at University of California, Berkeley, understands the appeal better than most people. In fact, it was a modern twist on a century-old pandemic that enticed him away from graduate studies in high-energy particle physics. About the time he was packing up to spend a year as a guest scientist at the Fermi National Accelerator Laboratory, he came across a story about how scientists were mapping mutations in the influenza virus from the 1918 flu pandemic to its molecular structure to explain how a bird virus became so virulent in people.
“I was absolutely captivated that a history-changing medical event could be explained using diffraction methods familiar from physics,” says Hurley. He cold-called one of the lead authors, the late Don Wiley at Harvard University, who encouraged him and suggested a number of places to apply for PhD programs in structural biology. Hurley chose Robert Stroud’s lab at University of California, San Francisco, to escort him into the world of membrane proteins. For postdoctoral work, he headed to Brian Matthews’ lab at University of Oregon and then spent 20 years at the NIH. He joined UC Berkeley in 2013.
The Hurley lab explores how proteins interact with membranes. They have a special interest in how proteins reshape phospholipid bilayers to sort cargo, direct metabolic processes, and create essential working structures within cells. They study how viruses coopt these activities to get in and out of cells, and how some defects in these processes lead to neurodegenerative diseases, such as Parkinson’s.
A strength of the Hurley lab is combining the structural biology with membrane reconstitution and membrane biophysics. “We’re trying to understand how the physical properties of membranes are modulated by proteins and how the modulation explains what happens in cell biology,” he says.
Along the way, the group has explored membrane interactions of other history-making viruses, HIV and, this year, SARS-CoV-2. For example, project scientist Xuefeng “Snow” Ren, postdoctoral fellow Cosmo Buffalo, and other Hurley lab colleagues found a structural explanation for how humans resisted infection by SIV, the ancestor of HIV, for millenia (Cell Host & Microbe, 2019).
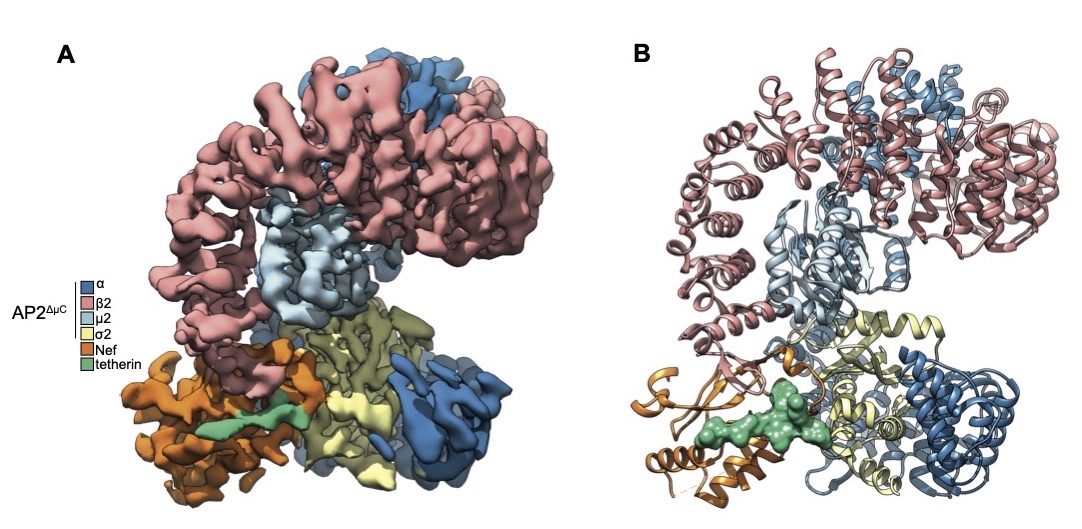
A cryo-EM density map shows how a part of the SIV virus called Nef disables a protein called tetherin that would otherwise block the virus from exiting the cell and mark it for destruction by the immune system. In people, the tetherin protein is missing the section that SIV Nef binds. The finding explains how humans resisted cross-species transmission of SIV from non-human primates. Image credit: Courtesy J.Hurley.
Then came the coronavirus, and the lab quickly mobilized its expertise around key questions related to how its proteins interact with host proteins and membranes, as well as collaborating with others on campus to apply this information to antiviral drug discovery. In the first project, one postdoctoral fellow caught in the pandemic lockdown on his way out of the lab was curious about why COVID-19 is worse than SARS. He and his fellow postdocs Buffalo and Rick Hooy determined the structure of the coronavirus’s ORF8 protein, a rapidly evolving protein thought to help evade the human immune system by attacking MHC-I, a tactic shared by HIV and many other viruses.
Hurley was born in Moscow, a college town in the Idaho panhandle. Restless
parents moved the family every year or two to wide ranging destinations,
including Vermont, Hawaii, Japan, Colorado, Pennsylvania, and California. As
soon as they arrived at their new home, family legend has it, his folks would
pull out the atlas and start thinking about where to go next.
Unbeknownst to him at the time, Hurley got a jumpstart on his future when he was 8 years old in Boulder, Colo. He started collecting minerals and learned the seven crystal systems—which describe the different symmetries of atoms—long before he ever heard of X-ray crystallography or struggled to persuade biological molecules to order themselves into one of these three-dimensional patterns for diffraction and analysis.
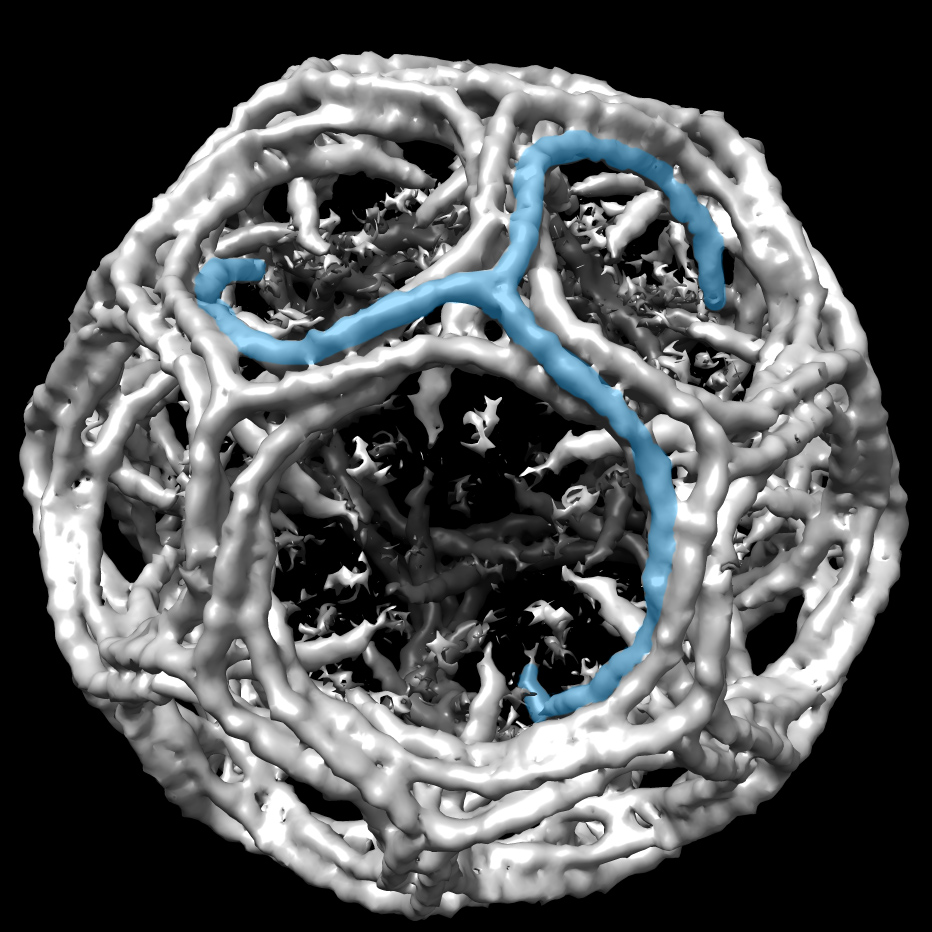
At the NIH, Hurley became interested in the ESCRT complexes. They prompt an unusual type of membrane budding that forms tiny structures that transport receptors and other membrane proteins to the lysosome. The buds form away from the cytosol instead of toward it, as with clathrin. And instead of severing the bud from the outside, ESCRT works from inside the neck, somehow not getting in the way of itself and its function.
“We have a model that I think makes sense,” Hurley says. “It’s very complicated and, at first glance, not terribly intuitive. I still would like to have a structural movie of the reaction. One of my remaining career goals is to really understand that.”
ESCRT influences a staggering number of functions, from fending off neurodegeneration to pruning neurites to pinching off the membrane neck between daughter cells after cell division, Hurley says. HIV coopts the ESCRTs to release newly made viruses.
In a 2018 Science paper, the lab built a new instrument to measure the force ESCRTs generate when severing experimental nanotubes. Hurley would like to be able to block the step when a virus hijacks the process, which could lead to a new class of antiretroviral therapeutics.
One of the lab’s more recent interests is autophagy, a way of recycling interior components of cells that seems bizarre at first glance, Hurley says. A double membrane sheet grows and engulfs the contents. (And these autophagosomes are closed off by ESCRTs).
In fact, the autophagy of mitochondria, or “mitophagy”, has become a central focus of the lab. In 2020, Hurley became a lead on a $7 million, six- lab grant on mitophagy.
“The biggest single project in my lab involves the self-digestion of cytosol,” he says. “It sounds like something that would be a bad idea, but it’s important to preserve healthy cells. It’s important for health of neurons to digest damaged mitochondria and digest molecular aggregates.”
An early finding detailed a complex important for organizing nucleation of the autophagosomal membrane. Autophagy is upregulated by diet and exercise and data suggests inducing it may be therapeutically helpful for incurable diseases. The work was spun out into a start-up as part of a UC Berkeley program that supports early-stage commercialization and eventually incorporated into another company called Casma Therapeutics in Cambridge, Mass., which is focusing on traditional small molecule drug discovery aimed at autophagy complexes.
As a teen, Hurley gravitated toward poetry and literature. But it wasn’t until he became a professional scientist that he published a poem. It was published in 2014 in the journal Autophagy, just as the lab was getting interested in the way a membrane cups around damaged mitochondria to remove and recycle them. The verses capture an imaginary conversation between a damaged mitochondrion and the autophagosome that intends to engulf it.
Hurley first wrote it to get a laugh in a seminar he was teaching. “Eyes glaze over after a huge amount of structural detail,” he says. “Sometimes I change gears by showing cell biology. Sometimes I do something pure fun.”
Hurley’s typical working day as a structural biologist at University of California, Berkeley, is bookmarked by a cycle through the hills above campus. During the pandemic restrictions, he still gets out his mountain bike, peddles one block to Tilden Park, and kicks up dust on the trails. But those days, the ride takes him back to his home office. Instead of going into the lab, Hurley gives his graduate students and postdoctoral fellows priority for the limited pandemic-rationed shifts there.
For fun, he hops on his mountain bike again three days a week in his role as a volunteer coach for the local composite middle school and high school cycling teams. He works with the entry level students to help them learn good judgment, techniques for safe descents, and general grit to keep going on cold and muddy days.
*Carol Cruzan Morton