Predicting Success
Bil Clemons
California Institute of Technology
Published December 18, 2018
For Bil Clemons, the first glimpse of a protein structure never gets old. “Seeing the electron density for the first time—it’s still just magical and one of the greatest experiences you can have as a scientist,” says Clemons, a biochemist at the California Institute of Technology (Caltech).
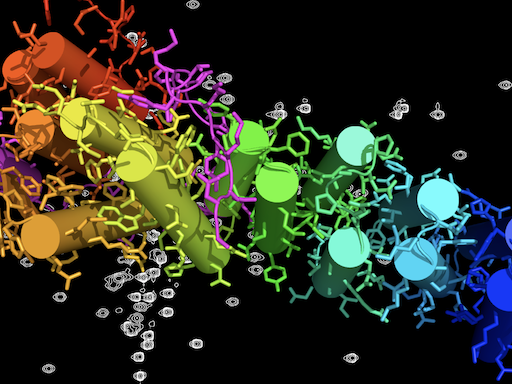

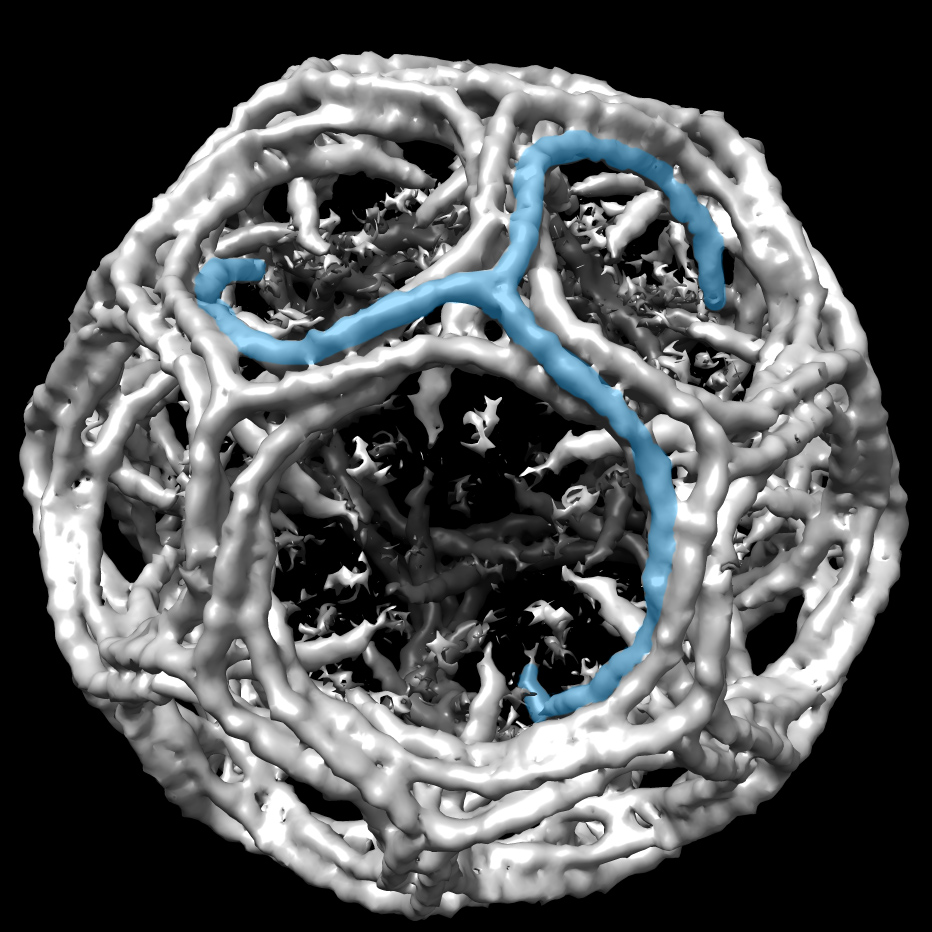
The crystal structure of Get4 bound to the N-domain of Get5 with the NMR spectra of the Get5 dimerization domain in the background. Image credit: Courtesy of B.Clemons
Those moments might be all the sweeter because of the months and years of trial and error that culminates in the final atomic image of a protein. As a graduate student, for example, Clemons put in 4 years to help solve the structure of the small ribosomal subunit, part of the molecular machine that translates genetic material into proteins in the cells. Now he works on labor-intensive membrane proteins.
Clemons got hooked on exploring the atomic terrain of molecules in part by a pair of 3D glasses when he was an undergraduate at Virginia Tech. The paper glasses were tucked into a special magazine on structural biology handed out in a class designed to help steer biochemistry students toward potential careers. Clemons ended up at the University of Utah in Salt Lake City. He was the first graduate student to join the lab of newly arrived Venki Ramakrishnan. Why did he make that choice? “I loved the concept of protein translation,” says Clemons.About four years into Clemons’ PhD, Ramakrishan moved the lab to the MRC Laboratory of Molecular Biology (LMB) in Cambridge, England, largely because of the stable funding for risky projects like the ribosome structure. Clemons made the transition and helped to finish the atomic resolution structure of the 30S subunit. In 2009, Ramakrishnan shared the Nobel Prize in Chemistry for the structure of the ribosome. Notably, Clemons features prominently in Ramakrishnan’s 2018 memoir of the path to the Nobel, “The Gene Machine.”
As a graduate student, Clemons weathered the tedious task of freezing hundreds of crystals one at a time in the cold room by playing music. Clemons, who always has music on in the background, went through a Johnny Cash phase as a graduate student. But truth be told, the saxophone is the family instrument.
Clemons’ father and namesake, William, was a U.S. Marine bandsman and sax player who grew up in a segregated south. His late uncle Clarence was a saxophonist with Bruce Springsteen’s E-Street Band, a gig subsequently inherited by Clemons’ younger brother Jake. Clemons’ mother was an elementary school teacher from North Dakota. Because of the military, the family moved around, but one constant influence on Bil was a passion for understanding the natural world exemplified by the availability of iconic shows such as ‘Mutual of Omaha’s Wild Kingdom’.
When he received his PhD, membrane proteins were becoming a feasible area of structural biology. Contemplating a postdoctoral fellowship, Clemons emailed Tom Rapoport at Harvard Medical School, a leader in membrane protein biogenesis, for advice on what group would be best for working on the structure of the protein translocation channel. Rapoport asked for his CV, then quickly invited Clemons to join his team working on the SecY channel, a team that included Stephen Harrison and members of his group.
At HMS, Clemons and his colleagues determined the structure of the universal channel that allows newly made proteins to be transported through or into cellular membranes. The finding was reported in Nature on New Year’s Day 2004.
“The ribosome was great, because no one had conquered it,” Clemons said. “But we knew what that mountain looked like. The structure didn’t change the general basics of what we knew. But with the protein translocation channel, the structure told us everything. Leveraging the known biochemistry with a structure, we could finally generate mechanistic models that could be clearly tested. It has stood the test of time.”
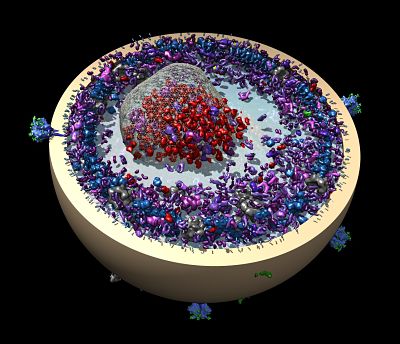
Membranes create protected spaces for living cells with a bilayer defining an inside and outside. Embedded membrane proteins span this barrier and provide a critical path for information and nutrients. A third of the genome encodes membrane proteins. Most membrane proteins are made next to the membrane, because their hydrophobic sections are incompatible with the watery environment inside the cell. To ensure correct insertion, for most membrane proteins, the ribosome cozies up to the endoplasmic reticulum and pushes the protein through the protein translocation channel, where the protein can be inserted into the membrane. There, it can fold and then, if necessary, be trafficked to various organelles.

At Caltech, Clemons has been studying another pathway used by a class of membrane proteins, known as tail-anchored proteins, to integrate into the membrane. “They have a special toolkit for getting into the lipid bilayer,” he says.
The lab has structurally characterized the various components that take the protein from the ribosome and deliver to the endoplasmic reticulum through the cytoplasm. “It’s a dance, a series of handoffs of the proteins, orchestrated player to player,” he says.
In the process, he has run the gamut of structural biology methods and pretty much all the software packages at SBGrid. But structure is just the starting point for figuring out the mechanism for the proteins in motion. His team probes the dynamics and chemistry using biochemical and biophysical tools “to understand at the deepest level how they’re working,” he says.
In addition to his work on membrane protein biogenesis, the lab has focused on other areas, such as the twin-arginine secretion pathway and membrane associated glycosylation pathways.
The Clemons lab has added another tool: Computational biology. They are trying to take some of the trial and error out of the difficult early stage of studying membrane proteins. “Caltech has enabled me to lean on students with a lot of computational training,” he says. “With that, we’ve tried to think about problems in our area that haven’t been addressed and try to solve them.”
A big problem for structural biologists begins at the first step. Scientists often insert the gene for protein of interest into certain bacteria or yeast, and then induce the host to generate the protein. For membrane proteins, this is a make-or-break first step, because “it is unlikely that one can get enough protein to study,” Clemons says. “Every other step in the process has a high failure rate, but if you don’t succeed in the first step, you will not succeed at all.”
He proposed using computers to design a statistical model that would predict how bacteria will react when asked to create a protein they don’t normally produce. With a key graduate student, his team showed that it is possible to predict membrane protein expression directly from sequence data, which allows researchers to double their odds of expressing a membrane protein. The findings were published in March 2018 in the Journal of Biological Chemistry.
“It’s early days,” Clemons says. “But just to demonstrate that we can leverage computation for this complicated problem is exciting for us. Large data sets, statistical tools and bioinformatics are opening up new paths. The synergy between computation and biochemistry is very powerful.”