Keep on Moving
James Berger
Johns Hopkins School of Medicine
Published August 23, 2015
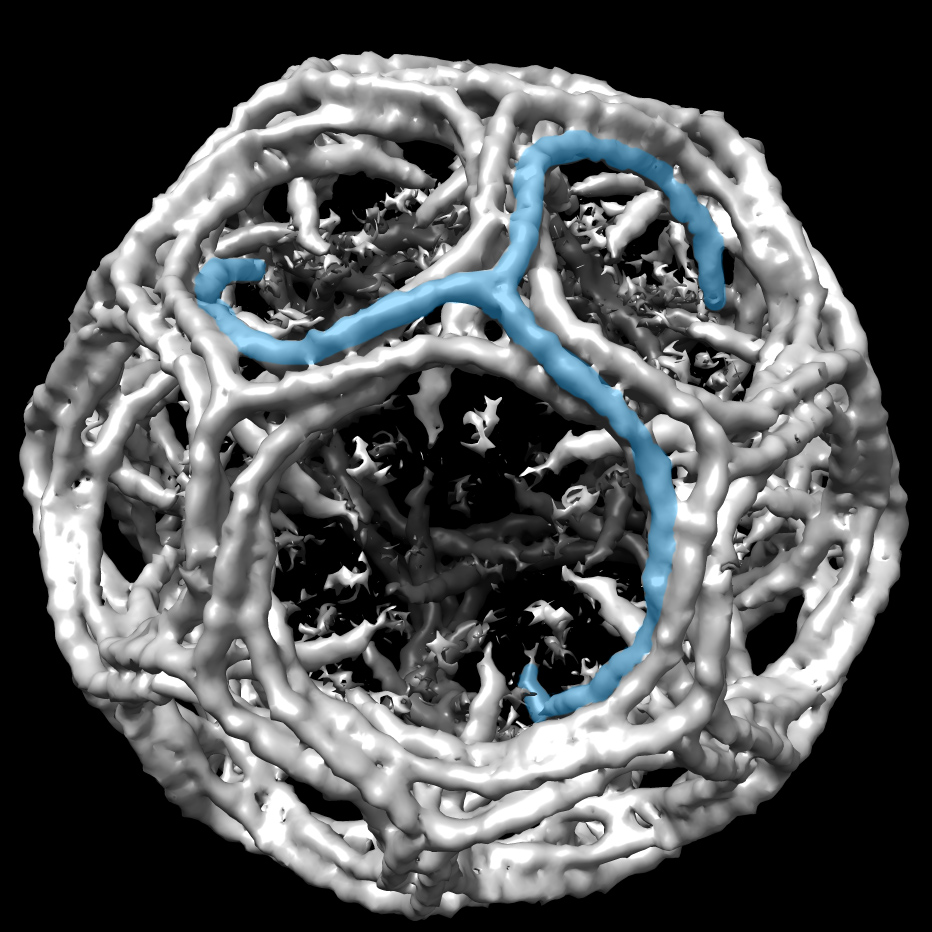
Say I’ve got two related motor proteins, says James Berger, professor of biophysics and biophysical chemistry at the Johns Hopkins School of Medicine, both helicases. One, which is responsible for DNA replication, moves at high speeds, up to 1000 base pairs per second. The other, involved in transcription, plods along at 30 bases per second. These proteins share a common ancestor, but have very different physical behaviors. “Why?” he asks. “What is it in the structure that encodes that maximum speed limit at which the motor can move? I would argue that you can look at any structure and you will have no idea.”
These are the questions Berger would like to spend all his time trying to answer. So far, structural biologists have done a good job of connecting structure to chemistry and chemical behaviors, but the gap between structure and physics remains wide. His lab’s efforts to close it will involve integrating structural biology with biophysical measures — and developing novel single molecule techniques in house to make these measures. “I think it will be really fun to try to figure out,” says Berger.
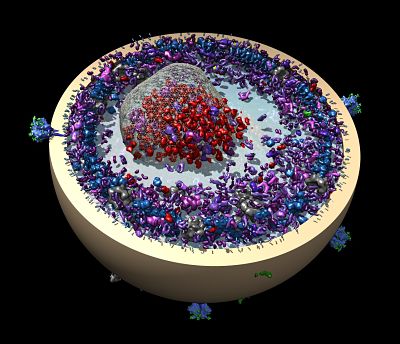
In reality, Berger’s lab also has many other projects to attend to. In a recent relocation from the University of California at Berkeley to Johns Hopkins, most of Berger’s lab moved with him and continues their work to understand the DNA replication system, both in terms of the larger more complex assemblies and in terms of the super-small interactions at the single molecule level. The move from a university to a medical school setting brings with it a slight change in perspective. Berger and his team now have more opportunities to think about the clinical applications of their work. “Partnering with clinicians here could lead us into new and interesting directions,” Berger says.
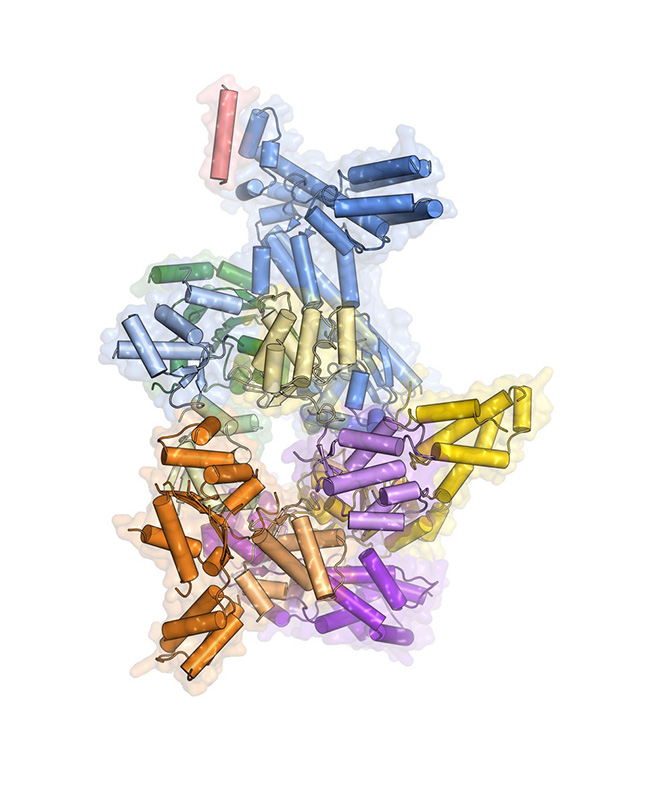
One of the latest crystal structures from the Berger lab shows how the the heterohexameric origin recognition complex (ORC), which is essential for coordinating the onset of DNA replication, can switch between two conformations, suggesting novel ways of controlling ORC functions.
Berger first found his way to structural biology as an undergraduate in biochemistry at the University of Utah. During the school year, he learned about proteins and biochemistry and in the summers he worked at Los Alamos National Laboratories with a researcher who was trying to understand how plants grow in contaminated soils. “I did an experiment with radio-labeled TNT once. It was diluted, but still,” says Berger. “These were things you wouldn’t see in a typical university lab.”
He went on to graduate school at Harvard University in 1990. He was interested in structural biology, but chose to work in the lab of biochemist James Wang. “Somehow it seemed you had to have the biochemistry lined up before you could do the structure,” says Berger. “I’d like to say it was foresight, but I suspect it was dumb luck.”Berger declared to Wang that he wanted to work on the structure of type II topoisomerases. Wang nodded and smiled. “I spent the next three years floundering,” Berger says.
Eventually, he determined the structure of the primary DNA cleaving region of yeast topo II. The accomplishment came with the help of Harvard structural biologist Stephen Harrison, who became Berger’s de-facto co-advisor. “He could point out folds just by looking at the structure. From memory,” says Berger. “Gaining insight into function just by looking at it was fun, and quite powerful.”
In 1995, Berger became a fellow at the Whitehead Institute at MIT and continued his work to understand motor proteins involved in packing and unpacking genes and winding and unwinding DNA. After his fellowship, he moved to Berkeley, where he began to focus on DNA replication. After several years of research, and several “aha” moments, Berger says, “we think we’re finally figuring out how this system works.”
Family needs triggered a move to Hopkins in 2013. While new projects focused on clinical applications have not yet begun, obvious possibilities include developing targets in the DNA replication system. This system is already targeted by anti-cancer drugs such as doxorubicin, and by antibacterials such as ciprofloxacin. “To me, the biggest existential threat is infectious disease. It pays to have this in the back of your mind,” says Berger. “How can we contribute to the process of drug discovery?”
In the realm of connecting structure to physics, the challenge, as in all of biology, is biological complexity. The number of possible structural variations is explosively large. “It’s complicated in a way that boggles the mind,” says Berger. “But the rules of chemistry and physics still apply. There’s power in that.”
-- Elizabeth Dougherty